Harnessing Agrobacterium for CRISPR: A Powerful Tool for Precision Plant Genome Engineering
This article provides a comprehensive guide for researchers and scientists on using Agrobacterium-mediated transformation for CRISPR/Cas delivery into plant cells.
Harnessing Agrobacterium for CRISPR: A Powerful Tool for Precision Plant Genome Engineering
Abstract
This article provides a comprehensive guide for researchers and scientists on using Agrobacterium-mediated transformation for CRISPR/Cas delivery into plant cells. We cover the foundational biology of Agrobacterium's gene transfer mechanism (T-DNA), detail step-by-step methodological protocols for vector design and plant transformation, and offer advanced troubleshooting for common challenges like low efficiency and off-target effects. The article also explores current validation techniques and comparative analyses with other delivery methods, highlighting Agrobacterium's unique advantages in generating stable, heritable edits for crop improvement and functional genomics.
The Engine of Change: Understanding Agrobacterium's Natural Gene Transfer for CRISPR
Abstract Agrobacterium tumefaciens has undergone a radical transformation from a studied plant pathogen to the preeminent vehicle for plant genetic engineering. This Application Note deconstructs its pathogenic machinery, reframes it as a programmable delivery system, and provides detailed protocols for its application in Agrobacterium-mediated CRISPR-Cas delivery to plant cells. The content supports a thesis on optimizing this system for high-efficiency, multiplexed genome editing in dicot and monocot species.
The Pathogenic Machinery: A Natural Genetic Engineer
The tumor-inducing (Ti) plasmid is the core pathogenic element. Key quantitative features of a representative nopaline-type Ti plasmid (pTiC58) are summarized below.
Table 1: Key Quantitative Features of the Ti Plasmid & T-DNA
| Component | Size / Location | Function |
|---|---|---|
| Total Ti Plasmid | ~200-250 kbp | Carries all essential virulence and T-DNA genes. |
| T-DNA Region | ~10-30 kbp (flanked by LB/RB) | Segment transferred and integrated into plant genome. |
| Left Border (LB) | 25 bp imperfect repeat | Recognition site for VirD2; less critical for transfer. |
| Right Border (RB) | 25 bp imperfect repeat | Primary initiation site for T-strand excision and transfer. |
| vir Region | ~40 kbp (7 major operons) | Encodes proteins for T-DNA processing, transfer, and regulation. |
Signaling & T-DNA Processing Pathway
Title: Agrobacterium Virulence Induction & T-complex Assembly
Protocol: Generating a CRISPR-Ready Agrobacterium Strain
Objective: To construct a disarmed Agrobacterium strain harboring a binary vector with a CRISPR-Cas9 expression cassette.
Materials (The Scientist's Toolkit): Table 2: Key Research Reagent Solutions
| Reagent/Material | Function/Description |
|---|---|
| Disarmed A. tumefaciens Strain (e.g., GV3101, LBA4404, AGL1) | Engineered with a "disarmed" Ti plasmid (vir genes present, T-DNA removed). Essential for transformation. |
| Binary Vector (T-DNA) | Plasmid containing LB/RB sequences flanking the gene(s) of interest (e.g., Cas9, gRNA, plant selectable marker). Replicates in both E. coli and Agrobacterium. |
| Electrocompetent Agrobacterium Cells | Cells prepared for transformation via electroporation. Higher efficiency than chemical methods. |
| SOC or LB Medium | Recovery medium post-electroporation. |
| Appropriate Antibiotics | Selection based on bacterial (e.g., rifampicin, gentamicin) and binary vector (e.g., kanamycin, spectinomycin) resistance markers. |
| Electroporator & Cuvettes (2mm gap) | For introducing DNA via electrical shock. |
| Liquid YEP or LB Media | For cultivation of Agrobacterium. |
Methodology:
- Binary Vector Preparation: Isolate high-purity plasmid DNA (e.g., using a mini-prep kit) from E. coli. Resuspend in sterile nuclease-free water or TE buffer. Concentration should be >100 ng/µL.
- Electroporation: a. Thaw 50 µL of electrocompetent Agrobacterium cells on ice. b. Add 50-100 ng of the binary vector plasmid DNA to the cells. Mix gently. Do not pipette vigorously. c. Transfer the mixture to a pre-chilled 2mm electroporation cuvette. Ensure no air bubbles are present. d. Pulse using an electroporator with settings: 2.5 kV, 25 µF, 400 Ω (for Agrobacterium; typical time constant ~4-5 ms). e. Immediately add 1 mL of pre-warmed (28°C) SOC or LB medium to the cuvette.
- Recovery & Selection: a. Transfer the cell suspension to a sterile 1.5 mL tube. Incubate horizontally at 28°C with shaking (200 rpm) for 2-4 hours. b. Plate 100-200 µL onto YEP or LB agar plates containing the appropriate antibiotics for the Agrobacterium chromosomal marker and the binary vector. c. Invert plates and incubate at 28°C for 48-72 hours until colonies appear.
- Confirmation: Pick 3-5 colonies, inoculate liquid cultures, and perform colony PCR or plasmid extraction followed by restriction digest to confirm the presence and integrity of the binary vector.
Protocol: Agrobacterium-mediated CRISPR Delivery to Plant Leaf Disks
Objective: To deliver a T-DNA encoding CRISPR-Cas9 components into plant cells via co-cultivation with Agrobacterium for stable transformation.
Experimental Workflow:
Title: Workflow for Agrobacterium CRISPR Delivery & Regeneration
Detailed Methodology:
- Plant Material Preparation: Surface-sterilize seeds of the target plant (e.g., Nicotiana benthamiana, Arabidopsis). Germinate on MS0 agar plates under sterile conditions. Use 3-4 week-old leaves.
- Agrobacterium Culture Induction: a. From a fresh plate, inoculate 5 mL of YEP/LB with antibiotics. Grow overnight at 28°C, 200 rpm. b. The next day, dilute the culture to OD600 = 0.1 in 10-20 mL of fresh medium (no antibiotics) containing 100-200 µM acetosyringone. c. Grow to OD600 = 0.5-1.0 (approx. 4-6 hours). Pellet cells at 4,000 x g for 10 min. d. Resuspend pellet in an equal volume of liquid plant co-cultivation medium (e.g., MS liquid + 100 µM acetosyringone).
- Excision & Inoculation: a. Using a sterile cork borer or scalpel, excise leaf disks (5-8 mm diameter) from the in vitro grown leaves. b. Immerse leaf disks in the prepared Agrobacterium suspension for 5-30 minutes with gentle agitation. c. Blot disks dry on sterile filter paper.
- Co-cultivation: a. Place disks abaxial side down on solid co-cultivation medium (e.g., MS + hormones + 100 µM acetosyringone). b. Seal plates with porous tape. Incubate in the dark at 22-25°C for 2-3 days.
- Selection & Regeneration: a. Transfer disks to selection/regeneration medium containing appropriate plant antibiotics (e.g., kanamycin, hygromycin) to inhibit Agrobacterium and select for transformed plant cells, and hormones (e.g., cytokinin for shoot induction). b. Subculture every 2 weeks to fresh selection medium. c. Once shoots develop (3-8 weeks), excise and transfer to root induction medium (e.g., MS + auxin).
- Analysis: a. After root development, transfer plantlets to soil. b. Extract genomic DNA from leaf tissue of putative transformants (T0). c. Screen for transgene presence via PCR for the Cas9 or selectable marker gene. d. Confirm CRISPR edits by sequencing the target genomic locus (e.g., using Sanger sequencing after PCR, or next-generation sequencing for multiplexed edits).
Quantitative Considerations for Optimization
Table 3: Key Optimization Parameters & Typical Ranges
| Parameter | Typical Range / Value | Impact on Efficiency |
|---|---|---|
| Bacterial OD600 at Co-cultivation | 0.5 - 1.0 | Higher OD can improve T-DNA delivery but increases overgrowth risk. |
| Co-cultivation Duration | 2 - 4 days | Longer periods increase transfer but also bacterial overgrowth. |
| Acetosyringone Concentration | 100 - 200 µM | Essential for vir gene induction in non-wounded setups. |
| Plant Tissue Age | 3-6 week leaves (species-dependent) | Younger, meristematic tissue generally more competent. |
| Selection Antibiotic Concentration | Species-specific (e.g., Kanamycin: 50-100 mg/L for tobacco) | Must be empirically determined to kill untransformed cells while allowing transformants to grow. |
| Agrobacterium Strain | GV3101, AGL1 (strong virulence) vs. LBA4404 | Affects host range and transformation efficiency. |
The efficacy of Agrobacterium tumefaciens-mediated transformation for delivering CRISPR-Cas components into plant cells hinges on the precise understanding and manipulation of the T-DNA transfer machinery. This system, encoded by the Ti (Tumor-inducing) plasmid, is a natural genetic engineer. For CRISPR delivery, the disarmed T-DNA region is engineered to carry expression cassettes for Cas nucleases and guide RNAs, flanked by the essential border sequences. The coordinated expression of the virulence (vir) genes, induced by plant phenolic signals, orchestrates the excision, transfer, and integration of this synthetic T-DNA. Optimizing this process for recalcitrant plant species or for achieving high-efficiency gene editing requires a detailed dissection of vir gene functions and border sequence interactions.
Core Components:virGenes and Borders
The Role of Border Sequences
The left border (LB) and right border (RB) are 25-bp direct repeats that delineate the T-DNA. The RB is critical, as the nicking event initiated by VirD2 occurs here to define the T-strand's 5' end.
Table 1: Characteristics of T-DNA Border Sequences
| Feature | Right Border (RB) | Left Border (LB) | Functional Implication |
|---|---|---|---|
| Core Sequence | 5'-TGACAGGATATATTGGCGGGTAAAC-3' | Similar 25-bp repeat | Recognition site for VirD1/VirD2 complex |
| Nicking Sites | Precisely between the 3rd and 4th base | Similar, but less efficient | RB nicking initiates T-strand production; LB nicks for release |
| Overdrive Sequence | Present ~25 bp upstream | Typically absent | Enhances VirD2 binding and T-strand efficiency |
| Role in CRISPR Delivery | Critical for T-DNA initiation; often the only border required in binary vectors | Can be omitted in "superbinary" or RB-only vectors; its presence may influence copy number | Simpler vectors with single RB can be effective for gene editing |
ThevirGene Regulon: Functions and Quantitative Expression
The vir genes (VirA, VirB, VirC, VirD, VirE, VirF, VirG, VirH) are located on the Ti plasmid and are essential for T-DNA processing and transfer. Their expression is tightly regulated by the VirA/VirG two-component system.
Table 2: Key vir Gene Operons and Their Functions in T-DNA Transfer
| vir Operon | Key Proteins | Primary Function | Optimal Induction Level (Relative Expression) | Relevance to CRISPR Delivery |
|---|---|---|---|---|
| virA/G | VirA (sensor kinase), VirG (response regulator) | Signal transduction; activates other vir genes in response to AS | VirG expression increases >50-fold upon induction | Target for constitutive activation (e.g., virG mutants) to bypass host-specific susceptibility. |
| virB | VirB1-B11 (Type IV Secretion System - T4SS) | Forms the transmembrane channel for T-complex & effector protein transfer | All 11 genes are co-expressed; essential for pore formation | Major host-range determinant; efficiency varies by Agrobacterium strain. |
| virC | VirC1, VirC2 | Binds Overdrive sequence to enhance T-strand production | Moderate inducer (5-10x increase) | Can be overexpressed to boost T-DNA excision in difficult plants. |
| virD | VirD1, VirD2, VirD4 | T-DNA processing; VirD1/D2 nick borders, VirD2 pilots T-strand | VirD2 is highly expressed post-induction | VirD2 nuclear localization signals (NLS) crucial for CRISPR cargo delivery to nucleus. |
| virE | VirE1 (chaperone), VirE2 (ssDNA-binding) | Coats T-strand in plant cell for protection & nuclear import | VirE2 is a major early-induced protein | Can be provided in trans from helper plasmid or engineered plant to widen host range. |
| virF | VirF (F-box protein) | Host protein degradation to facilitate integration | Low, host-dependent expression | May be omitted for monocot transformation; can affect regeneration. |
Application Notes for CRISPR Delivery Optimization
Note 1: Strain and Helper Plasmid Selection. For CRISPR delivery, use disarmed Agrobacterium strains (e.g., EHA105, GV3101) harboring a binary vector with CRISPR cassettes between the borders. Supplementing with a helper Ti plasmid containing extra copies of virG and virE (e.g., pTiBo542) can significantly enhance T-DNA transfer to monocots and recalcitrant dicots.
Note 2: Border Design in Binary Vectors. Modern binary vectors for CRISPR often use a single RB (RB-only vectors) to minimize transfer of vector backbone sequences, which is crucial for regulatory compliance. Ensure the RB sequence is intact and includes an overdrive enhancer for reliable nicking.
Note 3: Induction Conditions. Standard induction uses 100-200 µM acetosyringone (AS) in low-pH (5.2-5.6), sugar-rich medium at 22-25°C for 4-24 hours. Titration is critical: prolonged exposure to high AS can reduce Agrobacterium viability, while insufficient induction limits vir gene expression.
Table 3: Quantitative Effects of Induction Parameters on T-DNA Transfer Efficiency
| Parameter | Typical Optimal Range | Effect on vir Gene Expression | Measured Outcome on T-DNA Transfer |
|---|---|---|---|
| Acetosyringone [AS] | 100 - 200 µM | Maximal induction at ~200 µM; >500 µM can be inhibitory | Up to 10-fold increase in transient expression vs. no AS |
| Induction pH | 5.2 - 5.6 | virA activation optimal at pH <5.8 | Transfer efficiency can drop >50% at pH 7.0 |
| Induction Temperature | 22°C - 25°C | Optimal for protein stability/function; >28°C reduces efficiency | 25°C yields 2-3x more T-DNA complexes than 28°C |
| Induction Duration | 4 - 24 hours (strain dependent) | vir genes expressed within 2h; plateaus by 12-16h | Overnight (16h) induction standard for robust transfer |
Detailed Protocols
Protocol 1: AssessingvirGene Induction via RT-qPCR
Objective: Quantify the induction level of key vir genes (e.g., virD2, virE2, virG) under different conditions to optimize pre-induction for plant transformation. Materials: See "The Scientist's Toolkit" below. Procedure:
- Culture & Induce: Grow Agrobacterium harboring your binary + helper plasmids to mid-log phase (OD₆₀₀ ~0.6) in appropriate antibiotics. Pellet cells and resuspend in induction medium (e.g., MES buffer, pH 5.5, 2% sucrose, 200 µM AS). Incubate with shaking at 25°C for 0 (control) and 4-16 hours.
- RNA Extraction: Pellet 1-2 mL of induced culture. Use a commercial bacterial RNA extraction kit with on-column DNase I treatment to obtain high-purity RNA.
- cDNA Synthesis: Use 500 ng - 1 µg total RNA with a reverse transcription kit using random hexamers.
- qPCR Setup: Design primers for target vir genes and a stable reference gene (e.g., recA). Perform triplicate 10 µL reactions using SYBR Green master mix. Standard cycling conditions: 95°C for 3 min, followed by 40 cycles of 95°C for 10 sec, 60°C for 30 sec.
- Analysis: Calculate relative expression using the 2^(-ΔΔCt) method, comparing induced samples to the non-induced control.
Protocol 2: T-DNA Excision Assay (Border Nicking Assay)
Objective: Visually confirm the functionality of your border sequences and the VirD1/D2 complex by detecting the excised T-strand. Materials: See toolkit. Plasmid DNA containing your T-DNA with borders. Procedure:
- Provide VirD Proteins: Use an E. coli strain expressing VirD1 and VirD2 from an inducible plasmid OR use cell-free extract from induced Agrobacterium.
- In Vitro Nicking Reaction:
- Reaction Mix (50 µL): 500 ng supercoiled plasmid DNA, 25 mM MES-KOH (pH 5.6), 5 mM MgCl₂, 5 mM DTT, 2 mM ATP, and 10-20 µL of VirD-containing protein extract.
- Incubate at 25°C for 60 minutes.
- Stop Reaction: Add 1 µL of 10% SDS and 1 µL Proteinase K (20 mg/mL). Incubate at 37°C for 15 min.
- Analysis: Run the entire reaction on a 0.8% agarose gel in TAE buffer. Stain with ethidium bromide.
- Expected Results: Supercoiled plasmid (fastest migrating) will be converted to nicked open-circular (slower) and, if both borders nick, a linear T-strand fragment may be visible. Compare to negative control without protein extract.
The Scientist's Toolkit: Essential Research Reagents & Materials
Table 4: Key Reagents for Studying T-DNA Transfer Machinery
| Reagent / Material | Supplier Examples | Function in Protocol |
|---|---|---|
| Acetosyringone (AS) | Sigma-Aldrich, Thermo Fisher | Phenolic inducer of the vir gene regulon; critical for pre-induction. |
| Binary Vector Kit (e.g., pCAMBIA, pGreen) | Cambia, Addgene | Modular vectors with multiple cloning sites between LB/RB for CRISPR cargo. |
| Helper Ti Plasmid (e.g., pTiBo542) | Lab stocks, MO BIO | Provides supplemental vir genes to extend host range/transfer efficiency. |
| Supervirulent Agrobacterium Strains (EHA105, AGL1) | Lab stocks, ATCC | Disarmed strains with chromosomal backgrounds that enhance virulence. |
| MES Buffer | Sigma-Aldrich, Fisher BioReagents | Buffer for induction and transformation media; maintains optimal pH (~5.5). |
| Bacterial RNA Extraction Kit | Zymo Research, Qiagen | For high-quality RNA from Agrobacterium for vir gene expression analysis (RT-qPCR). |
| SYBR Green qPCR Master Mix | Bio-Rad, Thermo Fisher | For quantitative measurement of vir gene transcript levels. |
| Recombinant VirD1/VirD2 Proteins | Custom expression (e.g., Abcam services) | For in vitro border nicking/excision assays to validate T-DNA design. |
| Plant Tissue Culture Media | PhytoTech Labs, Duchefa | For co-cultivation and regeneration of transformed plant cells post Agrobacterium delivery. |
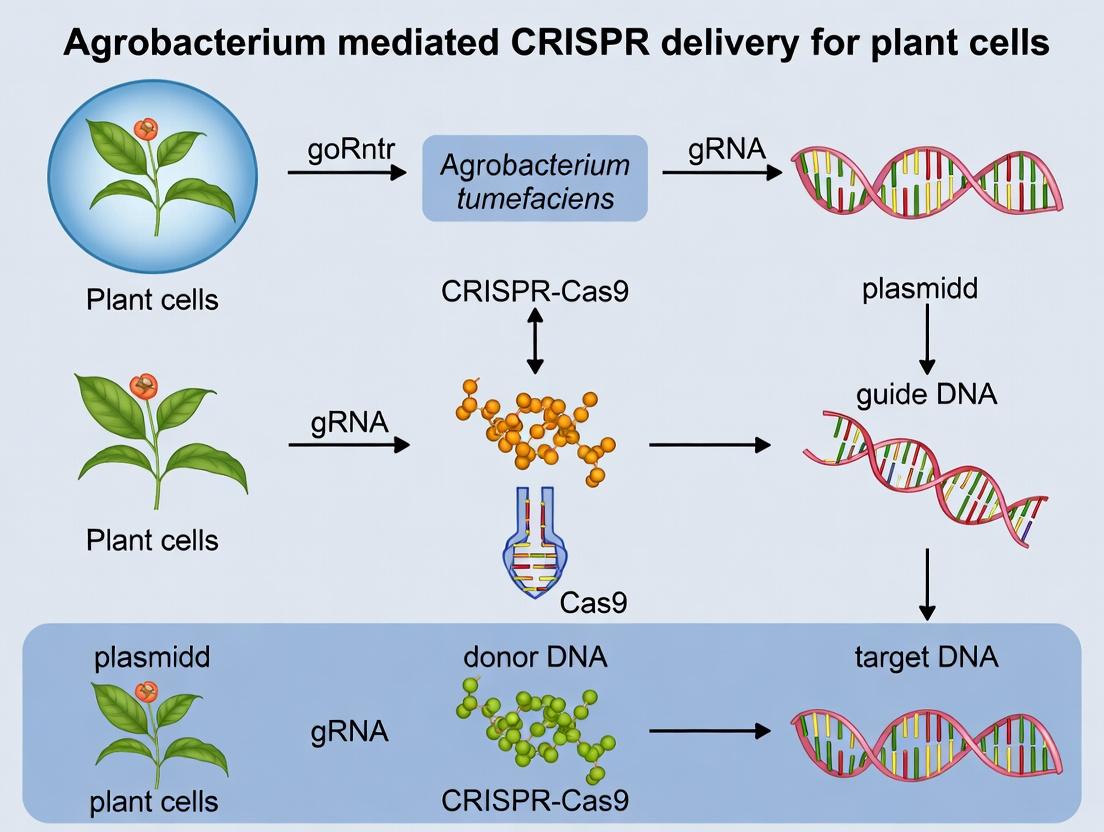
Within the context of Agrobacterium-mediated CRISPR delivery for plant cell research, the synergy between Agrobacterium tumefaciens and CRISPR/Cas systems represents a transformative approach. Agrobacterium naturally transfers DNA (T-DNA) into plant cells, a mechanism co-opted to deliver CRISPR components for precise genome editing. This application note details the rationale, protocols, and reagents for leveraging this synergistic technology, providing researchers with a robust framework for plant genome engineering.
The table below summarizes key quantitative advantages of using Agrobacterium over other common delivery methods for CRISPR in plants.
Table 1: Comparison of CRISPR Delivery Methods in Plants
| Parameter | Agrobacterium-mediated | PEG-mediated Protoplast Transfection | Biolistics (Gene Gun) |
|---|---|---|---|
| Transformation Efficiency | Moderate to High (varies by species) | Very High (in susceptible protoplasts) | Low to Moderate |
| Transgene Integration Rate | High (typically single-copy) | Very Low (mostly transient) | High (often complex, multi-copy) |
| Throughput | High (scalable for many explants) | Low (protoplast handling is delicate) | Moderate |
| Labor Intensity | Moderate | High | Low |
| Specialized Equipment Cost | Low | Moderate (electroporator) | Very High (gene gun) |
| Regeneration of Edited Plants | Direct from explants (e.g., leaf discs) | Requires protoplast-to-plant regeneration | Direct from tissues |
| Off-target Effects (Potential) | Lower (stable, low-copy integration) | Higher (transient, high copy number) | Higher (complex integration) |
| Applicability to Monocots | Improving with strain/supervisor optimization | High | Historically preferred |
Experimental Protocol: Agrobacterium-mediated CRISPR/Cas9 Delivery inNicotiana benthamiana(Leaf Disk Transformation)
This detailed protocol outlines the generation of stable gene-edited plants using a binary vector harboring CRISPR/Cas9 components.
Part A: Vector Construction and Agrobacterium Transformation
- Cloning: Assemble a sgRNA expression cassette (under a U6 or U3 Pol III promoter) and a Cas9 gene (under a 35S or ubiquitin promoter) into the T-DNA region of a binary vector (e.g., pCAMBIA1300, pGreenII). Include a plant selectable marker (e.g., hptII for hygromycin resistance).
- Agrobacterium Transformation: Introduce the recombinant binary vector into a disarmed, helper plasmid-containing Agrobacterium strain (e.g., EHA105, GV3101) via electroporation or freeze-thaw method.
- Detailed Method (Freeze-Thaw):
- Grow recipient Agrobacterium strain in 5 mL YEP (with appropriate antibiotics for helper plasmid) at 28°C to mid-log phase (OD₆₀₀ ~0.5-1.0).
- Chill cells on ice for 10 min, pellet at 4000 x g for 5 min at 4°C, and resuspend in 1 mL of 20 mM CaCl₂.
- Mix 100 µL of competent cells with 100-500 ng of plasmid DNA, freeze in liquid nitrogen for 5 min, and thaw at 37°C for 5 min.
- Add 1 mL of YEP broth, incubate at 28°C with shaking (220 rpm) for 2-4 hours.
- Plate on YEP agar with selective antibiotics for the binary vector and the helper plasmid. Incubate at 28°C for 2 days.
- Detailed Method (Freeze-Thaw):
Part B: Plant Transformation and Regeneration
- Pre-culture: Surface-sterilize N. benthamiana leaves and cut into 5x5 mm explants. Place explants on pre-culture MS medium (with hormones like cytokinin) for 2 days.
- Agrobacterium Co-cultivation:
- Inoculate a single colony of transformed Agrobacterium in 10 mL of induction medium (e.g., YEP with acetosyringone, typically 100-200 µM).
- Grow to OD₆₀₀ ~0.8. Pellet cells and resuspend in co-cultivation medium (liquid MS with acetosyringone) to OD₆₀₀ ~0.5.
- Immerse pre-cultured explants in the bacterial suspension for 10-30 minutes.
- Blot dry and transfer to solid co-cultivation medium. Incubate in the dark at 22-25°C for 2-3 days.
- Washing & Selection:
- Wash explants in sterile water or MS medium containing a bacteriostatic antibiotic (e.g., cefotaxime, 500 mg/L) to eliminate Agrobacterium.
- Transfer explants to selection/regeneration medium (MS with cytokinin, selective agent e.g., hygromycin, and bacteriostatic antibiotic).
- Regeneration: Transfer developing shoots to rooting medium (MS with auxin and selective agent). Once rooted, transfer plantlets to soil.
Part C: Molecular Analysis of Edited Plants
- Genomic DNA Extraction: Use a CTAB-based method from leaf tissue of putative transgenic plants.
- PCR Screening: Amplify the target genomic region and the transgene (e.g., Cas9, selectable marker).
- Mutation Detection: Subject PCR products to Sanger sequencing or use a restriction enzyme digest assay (if the CRISPR cut site disrupts a restriction site). For polyploid plants or complex edits, use next-generation sequencing (amplicon deep sequencing).
Visualizations
Title: Agrobacterium CRISPR Delivery Pathway
Title: Plant Transformation Workflow
The Scientist's Toolkit: Essential Research Reagent Solutions
Table 2: Key Reagents for Agrobacterium CRISPR Experiments
| Reagent/Material | Function/Purpose | Example/Notes |
|---|---|---|
| Disarmed Agrobacterium Strain | Engineered to transfer T-DNA without causing disease. Essential for stable plant transformation. | Strains: EHA105 (supervirulent), GV3101, LBA4404. Choice depends on plant species. |
| Binary Vector System | Carries CRISPR/Cas9 components within T-DNA borders for transfer into plant genome. | pGreen, pCAMBIA, pCAMBIA-UC series. Must be compatible with helper plasmid in strain. |
| Helper Plasmid | Provides vir genes in trans to mobilize T-DNA. Resident in many disarmed strains. | pSoup (in pGreen system), pTiBo542 (in EHA105). |
| Acetosyringone | Phenolic compound that induces the Agrobacterium vir gene system, critical for T-DNA transfer. | Add to both bacterial induction and co-cultivation media (100-200 µM). |
| Plant Tissue Culture Media | Supports growth, regeneration, and selection of transformed plant cells. | Murashige and Skoog (MS) basal medium, supplemented with hormones (e.g., BAP, NAA). |
| Selection Agents | Eliminates non-transformed plant tissue, allowing only edited cells to regenerate. | Hygromycin B, Kanamycin, Glufosinate. Agent depends on resistance gene in T-DNA. |
| Bacteriostatic Antibiotics | Eliminates residual Agrobacterium after co-cultivation without harming plant tissue. | Cefotaxime, Timentin (carbenicillin + ticarcillin). |
| PCR & Sequencing Reagents | For genotyping and confirming CRISPR-induced mutations at the target locus. | High-fidelity DNA polymerase for amplification, Sanger or NGS services for analysis. |
The broader thesis of this research program posits that Agrobacterium tumefaciens-mediated delivery of CRISPR-Cas components represents a transformative, broadly applicable platform for precise plant genome engineering. While historically robust in dicots, the recalcitrance of monocots to Agrobacterium infection has been a major bottleneck. This application note details the expanded host range of modern Agrobacterium-based vectors and provides updated protocols for key monocot families, directly supporting the thesis by enabling CRISPR-Cas delivery into previously challenging species. Success here is critical for translating genome editing technologies to staple cereals and bioenergy grasses.
Recent Advances in Host Range Expansion
The host range limitation has been overcome through synergistic optimizations in bacterial strain, vector design, and plant cell treatment.
- Super-Virulent Strains: The development of strains like AGL1 and LBA4404 Thy-, harboring extra copies of virulence (vir) genes or mutations to alleviate host-specific restrictions, has been pivotal.
- Ternary Vector Systems: The separation of T-DNA (carrying gene-of-interest and CRISPR machinery) and vir genes onto two different plasmids within the same bacterium enhances transformation efficiency and stability in monocots.
- Chemical Enhancers: The use of phenolic compounds (e.g., acetosyringone) to induce vir gene expression, and antioxidants (e.g., L-cysteine) to suppress plant defense responses during co-cultivation, is now standard.
- Tissue Culture Breakthroughs: Identification of highly regenerable explants, such as immature embryos in maize and wheat, and the use of morphogenic regulators (e.g., Baby boom / Wuschel2) to boost regeneration in non-embryogenic tissues.
Table 1: Transformation Efficiencies in Major Monocot Crops (2020-2024)
| Species | Common Name | Key Explant | Optimized Strain / System | Reported Efficiency (T0 Events) | Key CRISPR Application Demonstrated |
|---|---|---|---|---|---|
| Zea mays | Maize | Immature Embryo | AGL1 + Ternary Vector | 15-45% | Multiplexed editing for herbicide tolerance |
| Oryza sativa | Rice | Mature Seed Embryo | EHA105 | 40-85% | Biofortification (high iron/zinc) |
| Triticum aestivum | Wheat | Immature Embryo | AGL1 + VirG overexpression | 5-25% | Powdery mildew resistance (MLO knockouts) |
| Hordeum vulgare | Barley | Immature Embryo | AGL1 | 10-30% | Reduced dormancy & pre-harvest sprouting |
| Saccharum spp. | Sugarcane | Meristematic Buds | LBA4404 Thy- | 2-10% | Enhanced sucrose accumulation |
| Sorghum bicolor | Sorghum | Immature Embryo | AGL1 | 5-20% | Starch composition modification |
Detailed Experimental Protocols
Protocol A:Agrobacterium-Mediated Transformation of Immature Maize Embryos (B104 Inbred) for CRISPR Delivery
Objective: Generate stable, edited maize lines via Agrobacterium delivery of a CRISPR-Cas9 T-DNA.
I. Materials Preparation
- Plant Material: Immature ears harvested 10-12 days after pollination.
- Agrobacterium Strain: AGL1 harboring a ternary vector system (pVIR: vir genes; pCAS9: SpCas9; pTDNA: gRNA(s) and plant selection marker).
- Media: Infection Medium (IM), Co-cultivation Medium (COM), Resting Medium (RM), Selection Medium I & II (SM-I, SM-II). All contain 100 µM acetosyringone (IM, COM) or appropriate antibiotic/herbicide for selection.
- Enzymes & Chemicals: 0.1% Mercurochrome, 2% Sodium Hypochlorite, L-cysteine (added to IM/COM).
II. Step-by-Step Procedure
- Explant Isolation: Surface sterilize husked ears. Excise immature embryos (1.0-1.5 mm) into liquid IM.
- Agrobacterium Preparation: Grow AGL1 culture to OD600 ~0.8. Pellet and resuspend in IM + acetosringone + L-cysteine.
- Infection: Incubate embryos in Agrobacterium suspension for 10-15 min.
- Co-cultivation: Transfer embryos scutellum-side up on COM plates. Wrap and incubate in dark at 21°C for 3 days.
- Resting: Transfer embryos to RM plates (no selection, with Timentin to kill Agrobacterium). Incubate in dark at 28°C for 7 days.
- Selection I: Transfer developing calli to SM-I plates. Incubate at 28°C in low light for 14 days. Subculture surviving calli to fresh SM-I.
- Selection II & Regeneration: Transfer proliferating, resistant calli to SM-II (containing higher selection pressure and hormones to induce shoot formation). Subsequently transfer shoot primordia to regeneration medium for root development.
- Molecular Analysis: PCR-amplify target loci from putative transgenic plantlets for Sanger sequencing and Tracking of Indels by Decomposition (TIDE) analysis to confirm editing.
The Scientist's Toolkit: Research Reagent Solutions
| Reagent / Material | Function in Monocot Transformation | Example Product / Note |
|---|---|---|
| Super-virulent A. tumefaciens Strain AGL1 | Contains a disarmed C58 background & a super-virulent pTiBo542 plasmid. Broad host range, high efficiency in cereals. | Widely available from major culture collections. |
| Ternary Vector System Kit | Separates vir genes and T-DNA to reduce plasmid size, improve T-DNA delivery, and allow modular assembly of CRISPR components. | Kit systems from providers like Addgene or molecular biology suppliers. |
| Acetosyringone | Phenolic compound that induces the bacterial vir gene system, essential for initiating T-DNA transfer. | >98% purity recommended for reproducible results. |
| L-Cysteine (Antioxidant) | Suppresses plant tissue browning and hypersensitive cell death during co-cultivation, increasing viable explant count. | Filter-sterilize and add to media post-autoclaving. |
| Morphogenic Regulator Genes (Bbm/Wus2) | Transcription factors dramatically enhancing transformation and regeneration frequency in recalcitrant genotypes and explants. | Often included as a "helper" gene on the T-DNA or co-transformed. |
| Plant Preservative Mixture (PPM) | A broad-spectrum biocide/ fungicide used in tissue culture to suppress microbial contamination without harming plant tissues. | An alternative to traditional antibiotics like Timentin. |
Visualized Workflows & Pathways
Diagram 1: Agrobacterium-Monocot Interaction & CRISPR Delivery Workflow
Diagram 2: Ternary Vector System for CRISPR Delivery
From Plasmid to Plant: A Step-by-Step Protocol for CRISPR Delivery
Within the broader thesis on Agrobacterium-mediated CRISPR delivery for plant genome engineering, the design and assembly of the binary T-DNA vector is a critical foundational step. The efficiency of CRISPR-Cas9 editing in plant cells is directly influenced by the configuration of the expression cassettes for the Cas9 nuclease and the single-guide RNA (sgRNA). This protocol outlines current strategies for assembling these components into plant binary vectors, emphasizing modularity, efficiency, and suitability for Agrobacterium tumefaciens-mediated transformation.
Key Design Considerations and Quantitative Comparison
The choice of promoter, terminator, and assembly method impacts expression levels, specificity, and vector construction throughput. The following table summarizes quantitative data from recent studies (2023-2024) on key components.
Table 1: Comparison of Promoters for Cas9 and gRNA Expression in Plants
| Component | Promoter Name | Origin | Relative Expression Level* (Avg. % of Strongest) | Best Suited For | Key Reference(s) |
|---|---|---|---|---|---|
| Cas9 Nuclease | 2x35S (CaMV) | Virus | 100% (Baseline) | Dicots; High expression | LeBlanc et al. (2023) |
| ZmUbi1 | Maize | 95-110% | Monocots, Dicots | Wang et al. (2024) | |
| AtUBQ10 | Arabidopsis | 70-80% | Stable, constitutive expression | Silva et al. (2023) | |
| gRNA Scaffold | AtU6-26 | Arabidopsis | 100% (Baseline) | Dicots | LeBlanc et al. (2023) |
| OsU3 | Rice | 120-130% | Monocots | Chen et al. (2024) | |
| TaU6 | Wheat | 115-125% | Monocots, especially cereals | Park et al. (2023) |
*Expression levels are approximate and can vary based on plant species and tissue.
Table 2: Common Binary Vector Assembly Methods
| Method | Cloning Type | Typical # of gRNAs per Vector | Construction Time (Days) | Key Advantage | Throughput |
|---|---|---|---|---|---|
| Restriction Ligation | Standard | 1-2 | 5-7 | Low tech requirement | Low |
| Golden Gate | Type IIS | 4-10+ | 2-4 | High modularity & multiplexing | Very High |
| Gibson Assembly | Isothermal | 1-4 | 3-5 | Seamless, flexible | Medium |
| Gateway LR | Recombinational | 1-2 | 3-4 | Easy entry-to-destination cloning | Medium |
Detailed Protocols
Protocol 3.1: Golden Gate Assembly for Multiplex gRNA Expression Cassettes
This protocol describes assembling a polycistronic tRNA-gRNA array (PTA) into a binary vector containing a Cas9 expression cassette.
Materials:
- Enzymes: T4 DNA Ligase, BsaI-HFv2 (or Esp3I for MoClo).
- Vector: Binary backbone (e.g., pCambia, pGreen) with a plant selection marker and a Cas9 expression cassette (e.g., 2x35S::Cas9::NosT).
- Modules: Promoter (AtU6, OsU3) and tRNA-gRNA scaffold modules in Level 0 acceptor plasmids.
- Target Sequence Oligos: Designed 20-nt spacer oligonucleotides with appropriate overhangs.
Procedure:
- Design gRNA Spacers: Design 20-nt spacer sequences complementary to your target genomic loci. Verify specificity using tools like CRISPR-P 2.0 or CHOPCHOP.
- Anneal Oligos: Phosphorylate and anneal forward and reverse oligos to create a double-stranded spacer fragment with BsaI-compatible overhangs (4-bp stiches).
- Level 0 Assembly: Perform a Golden Gate reaction (37°C for 5 min, 16°C for 5 min, 20 cycles; then 50°C for 5 min, 80°C for 5 min) to clone each spacer into a universal gRNA scaffold module, creating Level 1 gRNA expression units (Promoter::gRNA::Terminator).
- Level 1 Assembly (Multiplexing): Assemble multiple Level 1 gRNA units, interspersed with tRNA sequences, into a single polycistronic array using a second Golden Gate reaction into an intermediate acceptor vector.
- Level 2 (Final) Assembly: Perform a final Golden Gate reaction to combine the polycistronic gRNA array from Step 4 with the binary destination vector containing the Cas9 cassette.
- Transformation & Verification: Transform the final reaction into E. coli, select on appropriate antibiotics, and verify constructs by colony PCR and Sanger sequencing across all assembly junctions.
Protocol 3.2:AgrobacteriumTransformation and Plant Cell Delivery
Materials: Electrocompetent Agrobacterium strain (e.g., GV3101, EHA105), binary vector plasmid, plant explants (e.g., leaf discs, callus).
Procedure:
- Electroporation: Introduce the verified binary vector into electrocompetent Agrobacterium cells. Recover cells in SOC/LB medium for 2-3 hours at 28°C.
- Selection & Culture: Plate on LB agar with antibiotics selective for the binary vector and the Agrobacterium strain (e.g., rifampicin, gentamycin). Incubate at 28°C for 2 days.
- Preparation for Co-cultivation: Inoculate a single colony into liquid induction medium (e.g., LB with appropriate antibiotics and 200 µM acetosyringone). Grow to OD600 ~0.8-1.0.
- Co-cultivation: Immerse sterilized plant explants in the Agrobacterium culture for 10-30 minutes. Blot dry and place on co-cultivation medium (with acetosyringone) for 2-3 days in the dark.
- Selection & Regeneration: Transfer explants to selection/regeneration medium containing antibiotics to kill Agrobacterium (e.g., cefotaxime) and select for transformed plant cells (e.g., kanamycin, hygromycin).
- Molecular Analysis: Extract genomic DNA from putative transgenic tissues. Confirm editing via PCR/RE assay (for indels) or sequencing of the target locus.
Visualizations
Title: CRISPR Binary Vector Assembly Strategy Flow
Title: Complete Experimental Workflow for Plant CRISPR
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for CRISPR Binary Vector Construction
| Reagent / Kit Name | Supplier Examples | Function in Protocol |
|---|---|---|
| BsaI-HFv2 & T4 DNA Ligase | NEB, Thermo Fisher | Core enzymes for Golden Gate assembly. BsaI cuts, and T4 Ligase joins fragments with compatible overhangs. |
| Gibson Assembly Master Mix | NEB | All-in-one enzyme mix for seamless, isothermal assembly of multiple DNA fragments. |
| Gateway LR Clonase II | Thermo Fisher | Enzyme mix for recombinational cloning of entry vectors into binary destination vectors. |
| pGreen/pCambia Vectors | Addgene, CABI | Standard binary vector backbones with T-DNA borders, plant selection markers, and bacterial origins. |
| MoClo Plant Toolkit Parts | Addgene | A standardized collection of Level 0, 1, and 2 plasmids (promoters, terminators, Cas9, gRNA scaffolds) for modular assembly. |
| Electrocompetent A. tumefaciens | Various Labs (GV3101) | Specialized strains optimized for plant transformation and vector maintenance. |
| Acetosyringone | Sigma-Aldrich | Phenolic compound that induces Agrobacterium's vir genes, essential for T-DNA transfer during co-cultivation. |
Within the broader research framework of Agrobacterium-mediated CRISPR delivery for plant genome editing, the preparation of robust, transformable Agrobacterium tumefaciens cultures is a foundational step. The efficiency of T-DNA transfer, carrying CRISPR-Cas9 components into plant cells, is directly influenced by the selected bacterial strain and the efficacy of transforming that strain with the desired binary vector. This application note details current best practices for strain selection and the creation of competent cells for high-efficiency transformation, critical for advancing plant functional genomics and trait development.
Agrobacterium Strain Selection for CRISPR Delivery
The choice of Agrobacterium strain is pivotal. Strains are differentiated by their chromosomal background and the presence of specific virulence (vir) genes. For CRISPR delivery, strains offering high transformation efficiency, minimal polysaccharide production, and compatibility with the plant species of interest are essential.
Table 1: CommonAgrobacterium tumefaciensStrains for Plant Transformation
| Strain | Chromosomal Background | Key Features & Suitability for CRISPR Delivery |
|---|---|---|
| GV3101 (pMP90) | C58 | Ti-plasmid replaced by disarmed helper plasmid pMP90. Excellent for Arabidopsis floral dip; moderate virulence. |
| LBA4404 | Ach5 | Carries helper Ti plasmid pAL4404. Widely used for monocots and dicots; relatively low virulence. |
| EHA105 | C58 | Derived from super-virulent A281, carries pTiBo542ΔT-DNA. High virulence, suitable for recalcitrant species. |
| AGL1 | C58 | Contains helper plasmid pTiBo542ΔT-DNA and a carbenicillin-resistance gene. High transformation efficiency, robust for many species. |
| C58C1 | C58 | Wild-type strain. Often used as a basis for engineered strains; requires a helper Ti plasmid for disarmed vectors. |
Current Trends (2023-2024): For CRISPR delivery, engineered strains like EHA105 and AGL1 are often preferred due to their high vir gene expression, which can improve T-DNA transfer of large CRISPR constructs. The use of strains with a C58 chromosomal background (e.g., GV3101, EHA105, AGL1) is recommended for most applications due to their robust growth and reliable virulence.
Protocol: Preparation of Chemically CompetentAgrobacteriumCells
This protocol describes a reliable method for generating chemically competent Agrobacterium cells suitable for high-efficiency transformation via freeze-thaw or electroporation.
Materials & Reagent Solutions
Table 2: Research Reagent Solutions for Competent Cell Preparation
| Reagent/Material | Function & Critical Notes |
|---|---|
| YEP Broth (Yeast Extract, Peptone) | Rich medium for vigorous growth of Agrobacterium to mid-log phase. |
| 10% (v/v) Glycerol Solution | Prepared in ultrapure water and filter-sterilized. Preserves cell viability during freezing. |
| 1 mM HEPES Buffer (pH 7.0) | Maintains a stable pH during cell washing to prevent stress. |
| 15% (v/v) Glycerol in YEP Broth | Freezing medium for long-term storage of competent cells at -80°C. |
| Liquid Nitrogen | For flash-freezing competent cell aliquots to maximize transformation efficiency. |
| Sterile Centrifuge Tubes | For pelleting bacterial cells without contamination. |
| Ice-cold Wash Buffers | All buffers must be pre-chilled to 0-4°C to slow bacterial metabolism. |
Detailed Protocol
Day 1: Inoculum Preparation
- Streak the selected Agrobacterium strain (e.g., AGL1) from a -80°C glycerol stock onto a YEP agar plate with appropriate antibiotics (if maintaining a helper Ti plasmid). Incubate at 28°C for 2 days.
Day 3: Culture for Competent Cells
- Pick a single, well-isolated colony and inoculate 5 mL of YEP broth (with antibiotics if needed). Incubate overnight (~16 hrs) at 28°C with vigorous shaking (250 rpm).
- The next morning, dilute the overnight culture 1:50 into 100 mL of fresh, pre-warmed YEP broth (without antibiotics) in a 500 mL flask.
- Incubate at 28°C with shaking (250 rpm) until the OD₆₀₀ reaches 0.5-0.6 (mid-log phase). This typically takes 4-6 hours. Monitor OD closely; overgrowth reduces competence.
Harvesting and Washing Cells
- Chill the culture on ice for 20 minutes. All subsequent steps must be performed in a cold room or on ice using pre-chilled solutions and tubes.
- Centrifuge the culture at 4,000 x g for 10 min at 4°C to pellet cells.
- Gently decant the supernatant. Resuspend the pellet completely in 40 mL of ice-cold, filter-sterilized 1 mM HEPES buffer (pH 7.0).
- Centrifuge again at 4,000 x g for 10 min at 4°C. Decant supernatant.
- Resuspend the pellet in 20 mL of ice-cold, filter-sterilized 10% glycerol solution.
- Centrifuge a third time at 4,000 x g for 10 min at 4°C. Decant supernatant carefully.
Final Resuspension and Aliquotting
- Gently resuspend the final cell pellet in 2 mL of ice-cold 15% glycerol in YEP broth. The suspension should be homogeneous.
- Immediately aliquot 50 µL or 100 µL volumes into sterile, pre-chilled microcentrifuge tubes.
- Flash-freeze the aliquots in liquid nitrogen and transfer to a -80°C freezer for long-term storage. Competent cells are now ready for transformation.
Protocol: Transformation of CompetentAgrobacterium(Freeze-Thaw Method)
This simple method is effective for introducing binary vectors (e.g., pCambia, pGreen series carrying CRISPR-Cas9) into competent Agrobacterium.
Procedure
- Thaw a 50 µL aliquot of competent cells on ice.
- Add 100-500 ng of plasmid DNA (binary vector) in a volume not exceeding 5 µL. Gently mix by flicking the tube. Do not vortex.
- Freeze the cell-DNA mixture in liquid nitrogen for 1 minute.
- Rapidly thaw the cells by placing the tube in a 37°C water bath for 5 minutes.
- Immediately add 500 µL of YEP broth (without antibiotics) to the tube.
- Incubate the tube at 28°C with shaking at 200 rpm for 2-4 hours for recovery and expression of antibiotic resistance genes.
- Plate 100-200 µL of the recovery culture onto YEP agar plates containing the selective antibiotics for both the Agrobacterium strain (e.g., rifampicin) and the binary vector (e.g., kanamycin, spectinomycin).
- Incubate plates at 28°C for 2-3 days until single colonies appear.
Key Workflow & Strain Selection Logic
Title: Agrobacterium Strain Selection Workflow for CRISPR
Binary Vector Transformation and Vir Gene Induction Pathway
Title: From Transformation to T-DNA Transfer Pathway
Application Note in the Context of Agrobacterium-mediated CRISPR Delivery
This protocol details a robust pipeline for Agrobacterium-mediated stable transformation of model dicot (e.g., Nicotiana benthamiana) and monocot (e.g., rice embryogenic callus) tissues, optimized for the delivery of CRISPR-Cas9 components. The procedure encompasses co-cultivation with engineered Agrobacterium, selection of transformed plant cells, and regeneration of whole plants. Success is critical for generating edited lines for functional genomics and trait development.
Key Research Reagent Solutions
| Reagent/Material | Function in Protocol |
|---|---|
| Agrobacterium tumefaciens Strain (e.g., EHA105, GV3101) | Disarmed Ti-plasmid harboring strain engineered to deliver T-DNA containing CRISPR-Cas9 expression cassettes into the plant genome. |
| Binary Vector with CRISPR Construct | Contains T-DNA borders flanking the gene of interest, Cas9 nuclease, and single-guide RNA (sgRNA) expression units, along with a plant selectable marker (e.g., hptII, bar). |
| Acetosyringone | Phenolic compound that induces Agrobacterium vir gene expression, enhancing T-DNA transfer efficiency during co-cultivation. |
| Plant Tissue Culture Media (MS, N6) | Basal nutrient media (Murashige and Skoog for dicots, N6 for monocots) supplemented with plant growth regulators (auxins, cytokinins) to induce callus or direct shoot regeneration. |
| Selection Antibiotic (e.g., Hygromycin, Kanamycin) | Selective agent added to culture media post-co-cultivation to inhibit the growth of non-transformed plant cells. Only cells expressing the transgenic resistance gene proliferate. |
| β-Lactam Antibiotic (e.g., Timentin, Carbenicillin) | Added to plant culture media after co-cultivation to eliminate residual Agrobacterium cells without harming plant tissues. |
Protocol 1: Co-cultivation with Agrobacterium
Objective: To facilitate T-DNA transfer from Agrobacterium harboring the CRISPR-Cas9 binary vector into plant explant cells.
Materials:
- Sterile explants (e.g., leaf discs, embryogenic callus)
- Agrobacterium culture (OD₆₀₀ = 0.5-0.8) in induction medium (containing 100-200 µM acetosyringone)
- Co-cultivation media (solid basal media + acetosyringone)
- Sterile blotting paper
Methodology:
- Explant Preparation: Aseptically prepare target tissues. For leaf discs, use a cork borer or scalpel.
- Bacterial Inoculation: Immerse explants in the Agrobacterium suspension for 5-30 minutes with gentle agitation.
- Co-cultivation: Blot explants dry on sterile paper and transfer to co-cultivation media. Incubate in the dark at 22-25°C for 2-3 days.
- Termination: Post incubation, transfer explants to recovery/selection media containing a β-lactam antibiotic to kill the Agrobacterium.
Table 1: Typical Co-cultivation Parameters for Different Explants
| Plant Species | Explant Type | Agrobacterium Strain | Co-cultivation Duration | Acetosyringone Conc. |
|---|---|---|---|---|
| Nicotiana benthamiana | Leaf Discs | GV3101 | 2 days | 200 µM |
| Oryza sativa (Rice) | Embryogenic Callus | EHA105 | 3 days | 100 µM |
| Arabidopsis thaliana | Floral Dip | GV3101 | N/A (In planta) | 0.05% Silwet L-77 |
Protocol 2: Selection of Transformed Plant Cells
Objective: To selectively promote the growth of plant cells that have integrated the T-DNA carrying the CRISPR-Cas9 and selectable marker genes.
Materials:
- Co-cultivated explants
- Selection media (basal media + plant growth regulators + selection antibiotic + β-lactam antibiotic)
Methodology:
- Recovery Phase (Optional): Transfer explants to media with a β-lactam antibiotic but without selection agent for 2-7 days to allow recovery and transgene expression.
- Primary Selection: Transfer explants to media containing the appropriate selection agent (e.g., 20-50 mg/L hygromycin). Subculture to fresh selection media every 2 weeks.
- Monitoring: Non-transformed tissues will bleach (chlorosis) and die over 2-4 weeks. Actively growing, healthy calli or shoots indicate putative transgenic events.
Table 2: Common Selection Agents and Concentrations
| Selectable Marker Gene | Selection Agent | Typical Working Concentration (mg/L) | Mode of Action |
|---|---|---|---|
| hptII (Hygromycin phosphotransferase) | Hygromycin B | 10-50 | Inhibits protein synthesis |
| nptII (Neomycin phosphotransferase) | Kanamycin | 50-100 | Inhibits protein synthesis |
| bar (Phosphinothricin acetyltransferase) | Phosphinothricin (PPT/Glufosinate) | 2-10 | Inhibits glutamine synthetase |
| epsps (Glyphosate resistance) | Glyphosate | 1-10 | Inhibits EPSP synthase |
Protocol 3: Regeneration of Whole Plants
Objective: To induce organogenesis or embryogenesis from selected transgenic calli to recover complete, edited plants.
Materials:
- Putative transgenic calli or shoots
- Shoot induction media (basal media + high cytokinin:auxin ratio)
- Root induction media (basal media + high auxin:cytokinin ratio or auxin only)
- Plant growth regulators (e.g., BAP, NAA, IAA, 2,4-D)
Methodology:
- Shoot Organogenesis: Transfer selected calli or tissue clusters to shoot induction media. Multiple shoot primordia should form within 2-4 weeks.
- Shoot Elongation: Excise developing shoots and transfer to media with a lower cytokinin concentration to promote elongation.
- Rooting: Transfer elongated shoots (≥1 cm) to root induction media. Roots typically develop within 1-3 weeks.
- Acclimatization: Once a healthy root system is established, gently wash off agar and transplant plantlets into sterile soil mix. Maintain under high humidity for 1-2 weeks before transferring to normal greenhouse conditions.
Table 3: Common Growth Regulator Regimes for Regeneration
| Species/Stage | Media Type | Cytokinin (e.g., BAP) | Auxin (e.g., NAA, IAA) | Other |
|---|---|---|---|---|
| Tobacco Shoot Induction | MS + 1.0 mg/L BAP | 1.0 mg/L | 0.1 mg/L NAA | - |
| Tobacco Rooting | ½ MS | - | 0.1-0.5 mg/L IAA | - |
| Rice Callus Induction | N6 + 2,4-D | - | 2.0 mg/L 2,4-D | - |
| Rice Shoot Regeneration | MS + BAP + NAA | 2.0-3.0 mg/L | 0.5-1.0 mg/L NAA | - |
Visualization of Protocols and Pathways
Agro-Transformation to Plant Workflow
Agro-CRISPR Delivery & Editing Pathway
Application Notes
Within the broader thesis on Agrobacterium-mediated CRISPR delivery for plant cells, these application notes detail three principal genome editing outcomes. The use of Agrobacterium tumefaciens for stable DNA integration remains a cornerstone for generating heritable edits in crops, enabling advanced functionalities beyond simple knockouts.
1. Gene Knockouts via Non-Homologous End Joining (NHEJ): This is the most common application for generating loss-of-function mutants. Agrobacterium delivers T-DNA carrying expression cassettes for CRISPR-Cas9 components. Double-strand breaks (DSBs) induced by Cas9 are repaired by the error-prone NHEJ pathway, often resulting in small insertions or deletions (indels) that disrupt the coding sequence. Success rates for generating biallelic knockouts in primary transformants (T0) for diploid crops typically range from 10-30%, depending on the target locus and transformation efficiency.
2. Base Editing: This application allows for precise, single-nucleotide changes without requiring a DSB or donor template. Agrobacterium delivers a base editor construct, which typically consists of a catalytically impaired Cas9 (nCas9 or dCas9) fused to a deaminase enzyme. Cytosine Base Editors (CBEs) enable C•G to T•A conversions, while Adenine Base Editors (ABEs) enable A•T to G•C conversions. Editing windows are typically 4-5 nucleotides within the protospacer. This is crucial for creating gain-of-function mutations or correcting deleterious SNPs. Efficiencies for desired base conversions in plants can exceed 50% in edited lines, with minimal indel byproducts (<5%).
3. Multiplexed Genome Editing: Delivering multiple single guide RNAs (sgRNAs) in a single T-DNA enables simultaneous editing of several loci. This is achieved through polymerase II or III-based expression of sgRNA arrays (e.g., tRNA-gRNA, Csy4-gRNA). Applications include knocking out redundant gene families, engineering complex metabolic pathways, and deleting large genomic segments. The main challenge is maintaining high editing efficiency for all targets, which often shows variance (e.g., 5-70% per target in the same construct).
Table 1: Comparison of Key Performance Metrics for CRISPR Applications in Crops via Agrobacterium
| Application | Typical Editing Efficiency (T0 Plants) | Primary Repair Pathway | Key Outcome | Indel Byproduct Frequency | Multiplexing Capacity (Common Range) |
|---|---|---|---|---|---|
| Gene Knockout | 10% - 80% (locus-dependent) | NHEJ | Frameshift mutations, premature stop codons | 100% (desired outcome) | 2-10 sgRNAs |
| Base Editing | 5% - 60% (base-dependent) | None (direct deamination) | Targeted point mutations | 0.1% - 10% | 1-5 sgRNAs |
| Multiplexed Editing | Varies per target (5-70%) | NHEJ or HDR | Multiple knockouts/edits | 100% for NHEJ targets | Up to 24+ sgRNAs (demonstrated) |
Table 2: Common Agrobacterium Vector Backbones and Selection Markers for Crop Editing
| Vector Backbone | Selection Agent (Plant) | Bacterial Selection | Typical Use Case |
|---|---|---|---|
| pCAMBIA2300 | Kanamycin (nptII) | Kanamycin | General CRISPR-Cas9 knockouts |
| pGreenII | Hygromycin (hptII) | Kanamycin | High-copy, multiplex gRNA constructs |
| pDIRECT | Glufosinate (bar) / Bialaphos | Spectinomycin | Cassette-free editing, base editing |
| pYLCRISPR | Hygromycin (hptII) | Spectinomycin | tRNA-gRNA multiplex systems |
Experimental Protocols
Protocol 1:Agrobacterium-Mediated Transformation for CRISPR Knockouts in Tomato Cotyledons
Objective: Generate stable, heritable gene knockouts in tomato (Solanum lycopersicum) cv. Micro-Tom.
Materials:
- Agrobacterium tumefaciens strain GV3101 carrying pCAMBIA2300-35S::Cas9-AtU6::sgRNA.
- Sterile tomato seeds.
- Co-cultivation Media (CM), Callus Induction Media (CIM), Shoot Induction Media (SIM), Root Induction Media (RIM) with appropriate hormones.
- Selection antibiotics: Kanamycin (100 mg/L for plants), Rifampicin (50 mg/L), and Carbenicillin (500 mg/L) for Agrobacterium.
Methodology:
- Vector Construction: Clone a 20-nt target-specific sequence into the sgRNA scaffold of the binary vector using BsaI golden gate assembly.
- Agrobacterium Preparation: Transform the binary vector into GV3101 via electroporation. Select single colony on YEP plates with Rifampicin and Kanamycin. Inoculate a 50 mL culture and grow to OD600 = 0.8. Pellet cells and resuspend in liquid CM + 200 µM acetosyringone.
- Explant Preparation & Infection: Surface-sterilize tomato seeds and germinate on hormone-free MS media. Excise cotyledons from 7-day-old seedlings and bisect. Immerse explants in the Agrobacterium suspension for 15 minutes, blot dry.
- Co-cultivation: Place explants on CM + acetosyringone in the dark at 25°C for 48 hours.
- Selection & Regeneration: Transfer explants to CIM + Kanamycin + Carbenicillin for 2 weeks to initiate callus. Subsequently, transfer to SIM + Kanamycin for shoot induction (3-4 weeks). Excise developing shoots and place on RIM + Kanamycin for root development.
- Molecular Analysis: Extract genomic DNA from putative transgenic plantlets (T0). Perform PCR amplification of the target region and subject to Sanger sequencing or next-generation sequencing (NGS) to detect indels. Use T7 Endonuclease I or Tracking of Indels by DEcomposition (TIDE) analysis for initial screening.
Protocol 2: Cytosine Base Editing in Rice Callus
Objective: Achieve C-to-T conversion in a target gene in rice (Oryza sativa) using Agrobacterium.
Materials:
- Agrobacterium strain EHA105 carrying pDIRECT_UBI::rBE3 (or similar CBE vector).
- Embryogenic rice calli (variety Nipponbare).
- N6-based transformation and selection media.
Methodology:
- Vector Design: Ensure the target C is within positions 4-8 (editing window) of the protospacer on the non-target strand for BE3 systems.
- Agrobacterium Co-cultivation: Follow standard rice transformation protocol. Mix freshly subcultured, embryogenic calli with the Agrobacterium suspension (OD600 = 0.1) for 30 minutes. Co-cultivate on filter papers over solid media for 3 days.
- Resting & Selection: Transfer calli to resting media with Carbenicillin (to kill Agrobacterium) for 5 days, then to selection media with Bialaphos for 2-3 weeks.
- Regeneration: Transfer resistant calli to pre-regeneration and then regeneration media to obtain plantlets.
- Analysis: Screen regenerated plantlets by sequencing the target locus. Quantify base editing efficiency via NGS of amplicons (recommended depth >10,000x). For a pure C-to-T change, screen for absence of indels by PCR/RE assay if the edit disrupts a restriction site.
Protocol 3: Multiplexed Gene Knockout using tRNA-gRNA System in Wheat
Objective: Simultaneously knockout three homologous TaMLO genes for powdery mildew resistance in hexaploid wheat.
Materials:
- Agrobacterium strain AGL1 carrying pYLCRISPR-Ubi::Cas9-AtU6::tRNA-gRNA array.
- Immature wheat embryos (variety Fielder).
Methodology:
- Construct Assembly: Synthesize or assemble a polycistronic tRNA-gRNA gene where each gRNA is flanked by endogenous tRNA sequences. Clone into a binary vector harboring Cas9.
- Wheat Transformation: Isolate immature embryos (1.0-1.5 mm). Infect with Agrobacterium suspension, co-cultivate for 2-3 days.
- Selection & Plant Regeneration: Transfer embryos to selection media with Hygromycin. Develop callus and regenerate plants over 12-16 weeks.
- Multiplex Screening: Isolate DNA and perform PCR across each target site. Use high-throughput sequencing (amplicon-seq) to deconvolute the complex indel patterns in all three homoeologs. Analyze for simultaneous biallelic/biallelic mutations.
Diagrams
Title: CRISPR-Cas9 Gene Knockout Workflow
Title: Cytosine Base Editing Mechanism
Title: Multiplex gRNA Vector Design
The Scientist's Toolkit
Table 3: Key Research Reagent Solutions for Agrobacterium-Mediated CRISPR in Crops
| Reagent/Material | Supplier Examples | Function in Application |
|---|---|---|
| CRISPR-Cas9 Binary Vectors (e.g., pCAMBIA, pYLCRISPR) | Addgene, Tsingke Biology | Plant transformation-ready plasmids containing Cas9, gRNA scaffold, and selection markers. |
| High-Efficiency Agrobacterium Strains (GV3101, EHA105, AGL1) | Lab stock, CIB | Engineered for superior T-DNA delivery to specific plant hosts (dicots/monocots). |
| Acetosyringone | Sigma-Aldrich | Phenolic compound that induces Agrobacterium vir gene expression, critical for T-DNA transfer. |
| Plant Tissue Culture Media (MS, N6, B5 basal salts) | Phytotech Labs, Duchefa | Provides nutrients and hormones for explant survival, callus induction, and plant regeneration. |
| Selection Antibiotics (Kanamycin, Hygromycin, Bialaphos) | GoldBio, Cell Signaling Tech | Selects for plant cells that have integrated the T-DNA carrying the corresponding resistance gene. |
| T7 Endonuclease I / Surveyor Nuclease | NEB, IDT | Detects CRISPR-induced indels by cleaving mismatched DNA heteroduplexes (initial screening). |
| Next-Generation Sequencing Kit (Amplicon-EZ) | Azenta, Illumina | Provides quantitative, high-depth sequencing of target loci to precisely characterize editing efficiency and patterns. |
| Uridine DNA Glycosylase (UDG) | Thermo Fisher | Used in BE4/4max systems to reduce unwanted C-to-T editing at off-target sites by degrading uracil. |
Maximizing Efficiency: Solving Common Problems in Agrobacterium-CRISPR Workflows
Within the broader thesis on optimizing Agrobacterium-mediated CRISPR-Cas9 delivery for plant genome editing, a primary bottleneck is low transformation efficiency. This inefficiency is a multifactorial problem rooted in the dynamic interplay between bacterial virulence and plant health/defense responses. This application note details the critical factors, diagnostic protocols, and reagent solutions for researchers to systematically identify and overcome these limitations.
Core Factors Impacting Transformation Efficiency
The success of T-DNA transfer and integration hinges on the precise coordination of bacterial virulence machinery and the susceptibility of the plant host. Disruptions in either domain dramatically reduce efficiency.
Bacterial Virulence Factors
Agrobacterium tumefaciens virulence is governed by a complex, induced genetic regulatory network.
Quantitative Data Summary: Key Virulence Genes and Their Impact
| Virulence Gene/Operon | Function | Reported Impact on T-DNA Transfer Efficiency (Knockout vs. Wild-Type) | Reference Strain |
|---|---|---|---|
| virA/virG | Two-component system; senses phenolics & sugars, activates vir regulon. | Reduction of 95-99% | LBA4404, EHA105 |
| virD1/virD2 | Endonuclease; processes T-DNA border sequences. | Reduction of >99% | GV3101 |
| virE2 | Single-stranded DNA-binding protein; coats T-strand for nuclear import & protection. | Reduction of 90-95% | AGL1 |
| virB Operon | Encodes Type IV Secretion System (T4SS) pilus. | Reduction of 98-99.9% | EHA105 |
| ChvE | Sugar-binding protein; augments virA sensitivity. | Reduction of 70-80% in certain hosts | C58 |
| AttM | Quorum-sensing lactonase; degrades autoinducer (3OC8-HSL). | Increase of 20-50% when attM is knocked out (prevents premature quenching) | C58 |
Plant Health & Defense Factors
The plant's physiological state and innate immune responses are equally critical.
Quantitative Data Summary: Plant Factors Influencing Susceptibility
| Plant Factor | Optimal Condition/Range for Transformation | Suboptimal Condition & Typical Efficiency Drop | Model Plant Data |
|---|---|---|---|
| Physiological Age | Exponential growth phase of explant cells. | Senescent tissue: Drop of 60-80% | Nicotiana tabacum leaf discs |
| Phenolic Production | Low basal levels, induced upon Agrobacterium perception (e.g., acetosyringone). | Constitutively high levels (stressed plants): Drop of 40-70% | Arabidopsis thaliana roots |
| Reactive Oxygen Species (ROS) | Tightly controlled, transient burst. | Chronic oxidative stress: Drop of 50-75% | Oryza sativa callus |
| Defense Hormones (e.g., SA, JA) | Low pre-induction levels. | Pre-activation of systemic acquired resistance (SAR): Drop of 60-90% | Solanum lycopersicum cotyledons |
| Cell Wall Composition | Actively dividing cells with "looser" walls. | Highly lignified/suberized cells: Drop of 70-95% | Zea mays immature embryos |
Diagnostic Protocols
Protocol 1: Assessing Bacterial Virulence Competence
Objective: To verify the induction and functionality of the Agrobacterium vir regulon. Materials: Induced bacterial culture, AS medium (pH 5.2-5.6), spectrophotometer, RT-PCR/qPCR reagents, reporter strain (e.g., harboring virB::GUS). Procedure:
- Culture & Induction: Grow Agrobacterium to mid-log phase (OD600 ~0.5-0.8) in minimal medium. Pellet and resuspend in induction medium (non-induced control: resuspend in minimal medium; induced sample: resuspend in medium + 200 µM acetosyringone). Incubate with shaking (200 rpm) at 25-28°C for 12-16 hours.
- Biomass Measurement: Record OD600 of induced vs. non-induced cultures. A slight reduction in growth of induced culture is normal due to metabolic burden.
- Molecular Verification (qPCR):
- Extract total RNA from 1 mL of control and induced cultures.
- Perform DNase treatment and cDNA synthesis.
- Run qPCR for a key virulence gene (e.g., virD2) and a housekeeping gene (e.g., recA).
- Calculate fold-induction using the 2^(-ΔΔCt) method. Expected induction: 50- to 200-fold.
- Reporter Assay (Optional): If using a reporter strain, perform a GUS/LacZ assay on induced cells and quantify activity spectrophotometrically.
Protocol 2: Evaluating Plant Explant Health and Defense Status
Objective: To quantify pre-transformation stress and defense markers in plant explants. Materials: Target plant explants, liquid N2, mortar & pestle, assay kits for H2O2, phenolics, antioxidants (e.g., APX, CAT). Procedure:
- Explant Preparation: Surface-sterilize and prepare explants (e.g., leaf discs, callus) as per standard protocol. Divide into batches.
- Biomarker Extraction: Flash-freeze 100 mg of explant tissue in liquid N2. Homogenize and extract in appropriate buffers (e.g., phosphate buffer for ROS assays, methanol/acetone for phenolics).
- Quantitative Assays:
- H2O2 Content: Use a colorimetric peroxide assay kit. Optimal baseline: < 5 µmol/g FW. High stress: > 15 µmol/g FW.
- Total Phenolic Content: Use the Folin-Ciocalteu method. Optimal baseline: Host-specific, but a 2-3 fold increase over healthy control indicates stress.
- Antioxidant Enzyme Activity: Measure catalase (CAT) and ascorbate peroxidase (APX) activity kinetically. Chronically high activity may indicate sustained oxidative stress.
- Interpretation: Explants showing high baseline H2O2, elevated phenolics, and high antioxidant activity are likely under stress and will exhibit lower susceptibility.
The Scientist's Toolkit: Key Research Reagent Solutions
| Reagent / Material | Function & Application | Key Consideration |
|---|---|---|
| Acetosyringone (AS) | Phenolic signal molecule; induces the Agrobacterium vir regulon. Critical for most non-Nicotiana hosts. | Use high-purity DMSO stock (e.g., 100 mM). Optimal co-culture concentration: 100-200 µM. Light-sensitive. |
| L-Cysteine / Dithiothreitol (DTT) | Anti-browning/anti-oxidant agents. Added to co-culture or pre-culture media to scavenge ROS, reduce explant necrosis. | Concentration is critical (e.g., 400 mg/L L-Cys). Too high can be bacteriostatic. |
| Silwet L-77 | Non-ionic surfactant. Enhances Agrobacterium adherence and infiltration into plant tissues. | Use at very low concentrations (0.005-0.02%). Can be phytotoxic; requires optimization. |
| Plant Preservative Mixture (PPM) | Broad-spectrum biocide. Used in co-culture to suppress Agrobacterium overgrowth without common antibiotics that stress plant cells. | Allows longer co-culture without bacterial overgrowth, improving transformation in sensitive explants. |
| DPI (Diphenyleneiodonium chloride) | Inhibitor of NADPH oxidase, a key enzyme in plant ROS burst. Can be used in diagnostic assays to quell defense response. | Tool for testing if ROS burst is the primary cause of low efficiency. Use in pre-treatment media. |
| Competent E. coli (e.g., Stbl3) | For stable maintenance of CRISPR binary vectors with direct repeats. Reduces risk of T-DNA rearrangement. | Essential for cloning and propagating CRISPR-Cas9 constructs prior to Agrobacterium electroporation. |
Visualization: Pathways and Workflows
Application Notes
Within the framework of Agrobacterium-mediated CRISPR delivery for plant genome engineering, a primary challenge is the frequent silencing of transgenes, including the Cas9/gRNA expression cassette. This silencing, driven by the plant's innate defense against foreign DNA, leads to unstable and non-persistent editing activity. Key strategies to overcome this involve the use of regulatory sequences and genetic elements that evade or suppress silencing mechanisms.
Table 1: Strategies to Mitigate Transgene Silencing in Plants
| Strategy | Mechanism | Typical Efficacy (Relative Expression Increase) | Key Considerations |
|---|---|---|---|
| Intron-Containing Cas9 | Spliceosomal activity disrupts silencing RNA signals, enhances mRNA processing. | 2- to 10-fold | Position and number of introns are critical. |
| Matrix Attachment Regions (MARs) | Insulate transgene from positional effects, may create independent chromatin domains. | 1.5- to 5-fold | Effect is sequence- and genomic locus-dependent. |
| Silencing Suppressor Proteins | e.g., P19, HC-Pro. Actively inhibit RNAi pathways that mediate silencing. | 5- to 50-fold | Risk of pleiotropic effects on plant development. |
| Epigenetically Neutral Promoters | Use of specific promoters (e.g., Z. mays Ubi, Cestrum YLCV) less prone to DNA methylation. | 3- to 8-fold | May have species-specific activity patterns. |
| CRISPR/Cas Transcriptional Fusions | Fusion to plant-derived ORFs (e.g., ubiquitin) to mimic endogenous transcripts. | 2- to 6-fold | Must ensure fusion does not impair Cas9 protein function. |
Recent data indicates that combining intron-optimized codon usage for Cas9 with MAR flanking sequences can yield a synergistic effect, improving stable expression rates in T1 lines from <20% to >70%.
Protocols
Protocol 1: Assembly of an Intron-Enhanced Cas9 Expression Cassette for Agrobacterium Vectors
Objective: To clone a plant-optimized Cas9 gene containing a plant-specific intron into a binary vector for stable transformation.
Materials:
- pRGEB32-like binary vector (with gRNA scaffold).
- Cas9 gene fragment with soybean FAD2 intron (or similar) inserted at a conserved location.
- Strong plant promoter (e.g., ZmUbi, CaMV 35S enhancer tandem).
- NOS or other polyA terminator.
- Gateway BP/LR Clonase II or standard restriction enzymes (e.g., BsaI, Golden Gate assembly).
- Chemically competent E. coli and Agrobacterium tumefaciens strain EHA105/AGL1.
Procedure:
- In-silico Design: Identify a permissive site in the Cas9 coding sequence (often within the first 300 bp) for intron insertion. Ensure the intron maintains the reading frame.
- Fragment Assembly: Using Golden Gate or Gibson assembly, combine the promoter, intron-containing Cas9 CDS, and terminator. This forms the "Expression Cassette."
- Binary Vector Assembly: Clone the assembled Expression Cassette into the T-DNA region of the binary vector, upstream of the gRNA expression unit(s).
- Validation: Sequence the final construct across all assembly junctions and the intron region.
- Transformation: Electroporate the validated plasmid into competent Agrobacterium cells. Select on appropriate antibiotics (e.g., rifampicin, spectinomycin).
Protocol 2: Co-delivery of a Silencing Suppressor for Transient Expression Boost
Objective: To transiently express the Tomato bushy stunt virus P19 protein alongside CRISPR components to maximize initial editing efficiency before silencing occurs.
Materials:
- Agrobacterium strain harboring the CRISPR binary vector.
- Agrobacterium strain harboring a P19 expression vector (e.g., pTKB-P19).
- Induction Medium (IM) with acetosyringone.
- Target plant tissue (e.g., Nicotiana benthamiana leaves, plant explants).
Procedure:
- Culture Preparation: Grow both Agrobacterium cultures overnight. Centrifuge and resuspend to an OD600 of ~0.5 in IM with 200 µM acetosyringone. Incubate 2-4 hours.
- Mixed Infiltration: Mix the CRISPR and P19 strains at a 1:1 ratio. For leaf infiltration, use a syringe to pressure-infiltrate the mix into the abaxial side of leaves.
- Co-cultivation: For explants, immerse them in the mixed bacterial suspension for 10-30 minutes, blot dry, and co-cultivate on solid medium for 2-3 days.
- Analysis: Assess Cas9 expression and editing efficiency 3-7 days post-infiltration via fluorescence (if tagged), Western blot, and T7E1 or sequencing assays. Note: This is for transient assessment; P19 is not intended for stable transformation.
Visualizations
Title: Workflow for Stable CRISPR Expression in Plants
Title: Silencing Pathways and Engineering Countermeasures
The Scientist's Toolkit
Table 2: Essential Research Reagent Solutions
| Item | Function in Experiment | Key Consideration |
|---|---|---|
| Binary Vector (e.g., pRGEB32, pYLCRISPR) | Carries T-DNA with CRISPR/Cas and selection marker for Agrobacterium-mediated delivery. | Choose modular systems (Golden Gate, Gateway) for easy part swapping. |
| Intron-Containing Plant Codon-Optimized Cas9 | Enhances mRNA stability and nuclear export, evading PTGS. Increases expression levels. | Verify intron is correctly spliced in your plant species. |
| Matrix Attachment Region (MAR) Sequences | Flank T-DNA to insulate from repressive chromatin, reducing positional silencing. | Effects can be variable; empirical testing is needed. |
| Silencing Suppressor Strain (e.g., AGL1+pTKB-P19) | Used for transient assays to maximize short-term expression and editing efficiency. | Not for generating stable transgenic plants due to developmental defects. |
| Acetosyringone | Phenolic compound that induces Agrobacterium vir gene expression, critical for T-DNA transfer. | Use fresh stock solutions; optimal concentration varies by plant species/tissue. |
| Plant Tissue Culture Media (e.g., MS, B5) | For selection and regeneration of transformed plant cells after co-cultivation with Agrobacterium. | Hormone composition (auxin/cytokinin ratio) is species- and tissue-specific. |
| CRISPR Editing Efficiency Assay Kit (e.g., T7E1, GUIDE-seq) | To quantitatively measure mutation induction frequency post-transformation. | Next-generation sequencing provides the most accurate and comprehensive data. |
Strategies to Minimize CRISPR Off-Target Effects in Plant Genomes
Within the context of Agrobacterium-mediated CRISPR/Cas delivery for plant genome engineering, off-target effects remain a significant hurdle to achieving precise, predictable modifications. These unintended edits can confound phenotypic analysis and raise regulatory concerns. This document outlines current strategies and provides detailed protocols to enhance on-target specificity in plant systems.
Table 1: Summary of Strategies to Minimize Off-Target Effects
| Strategy Category | Specific Method/Tool | Typical Reduction in Off-Target Effects (Quantitative Data) | Key Considerations for Agrobacterium Delivery |
|---|---|---|---|
| Cas Protein Engineering | High-Fidelity Cas9 variants (e.g., SpCas9-HF1, eSpCas9) | 60-95% reduction in detectable off-target sites (based on mammalian/plant cell studies) | Codon optimization for plants is essential; ensure expression cassette fits T-DNA. |
| gRNA Design Optimization | Use of predictive algorithms (CRISPR-P, CHOPCHOP) & truncated gRNAs (tru-gRNAs, 17-18nt) | tru-gRNAs can reduce off-target activity by >5000-fold while retaining on-target efficiency. | gRNA sequence must be cloned into appropriate Agrobacterium binary vectors. |
| Ribonucleoprotein (RNP) Delivery | Direct delivery of pre-assembled Cas9 protein-gRNA complexes | Off-target effects significantly lower than plasmid-based delivery; up to ~90% reduction. | Requires biolistic or protoplast transfection; not directly compatible with standard Agrobacterium T-DNA. |
| Anti-CRISPR Proteins | Transient expression of AcrIIA4/AcrIIA2 to inhibit Cas9 after editing window | Can suppress off-target activity by >90% while preserving on-target edits. | Requires a second T-DNA or co-infiltration strategy; timing of expression is critical. |
| Dual Nickase Strategy | Use of paired Cas9 D10A nickases with offset gRNAs | Can reduce off-target mutations to nearly undetectable levels (~10- to 150-fold reduction). | Requires two specific gRNAs to be expressed from the same T-DNA; larger T-DNA. |
| Promoter Selection | Use of cell type-specific or inducible promoters (e.g., heat-shock, estrogen-inducible) | Limits Cas9 expression temporally/spatially, reducing overall off-target potential. | Compatible with Agrobacterium; inducible systems require careful protocol optimization. |
Detailed Application Notes and Protocols
Protocol 1:Agrobacterium-Mediated Delivery of High-Fidelity Cas9 forNicotiana benthamianaLeaves
Objective: To achieve high-specificity genome editing using SpCas9-HF1 delivered via Agrobacterium tumefaciens. Research Reagent Solutions:
- pTRANS_SpCas9-HF1: Binary vector containing plant codon-optimized SpCas9-HF1 driven by a 35S promoter.
- pTRANS_gRNA: Modular binary vector containing the target-specific gRNA under a U6/U3 promoter.
- GV3101(pSoup) Agrobacterium Strain: Optimized for plant transformation, containing helper plasmid for T-DNA transfer.
- Acetosyringone Solution (100 mM): Phenolic inducer of Agrobacterium vir genes, essential for T-DNA transfer.
- LB Medium with Appropriate Antibiotics: For selection of binary vectors (e.g., Spectinomycin, Kanamycin).
- MS Infiltration Buffer (10 mM MgCl₂, 10 mM MES, 150 µM Acetosyringone, pH 5.6): Buffer for resuspending bacteria for leaf infiltration.
Methodology:
- Vector Construction: Clone the target-specific 20nt spacer sequence into the BsaI site of the pTRANS_gRNA vector. Verify by sequencing.
- Transformation: Co-transform pTRANSSpCas9-HF1 and pTRANSgRNA into competent A. tumefaciens strain GV3101(pSoup) via electroporation.
- Culture Preparation: Inoculate single colonies in 5 mL LB with antibiotics. Grow overnight at 28°C, 200 rpm. Sub-culture 1:50 into fresh medium with antibiotics and 20 µM acetosyringone. Grow to OD₆₀₀ ~0.8.
- Bacterial Preparation: Pellet cells at 3000 x g for 15 min. Resuspend pellet in MS Infiltration Buffer to a final OD₆₀₀ of 0.5. Incubate at room temperature for 2-4 hours.
- Plant Infiltration: Using a needleless syringe, infiltrate the bacterial suspension into the abaxial side of 4-week-old N. benthamiana leaves.
- Sampling and Analysis: Harvest leaf discs at 3-5 days post-infiltration. Extract genomic DNA and perform targeted deep sequencing (e.g., amplicon sequencing) of the on-target and predicted top off-target loci to assess editing specificity.
Protocol 2: Dual Nickase Strategy for Rice Protoplasts (Pre-Testing forAgrobacteriumDelivery)
Objective: To pre-validate the high-specificity editing of a target locus using Cas9 D10A nickase pairs in a transient protoplast system, prior to stable transformation via Agrobacterium. Research Reagent Solutions:
- pUbi-Cas9D10A-Nickase Plasmid: Expresses plant codon-optimized Cas9 D10A nickase under a maize ubiquitin promoter.
- pU6-gRNA Expression Vectors: Two vectors expressing offset gRNAs (spacing 20-70 bp) targeting opposite DNA strands.
- Rice Protoplast Isolation Enzyme Solution (1.5% Cellulase R10, 0.75% Macerozyme R10 in 0.4M Mannitol, pH 5.7).
- PEG-Calcium Transformation Solution (40% PEG4000, 0.2M Mannitol, 0.1M CaCl₂).
- W5 Solution (154mM NaCl, 125mM CaCl₂, 5mM KCl, 2mM MES, pH 5.7).
Methodology:
- Protoplast Isolation: Slice 2-week-old rice seedling stems into 0.5mm strips. Digest in 10 mL enzyme solution for 6 hours in the dark with gentle shaking. Filter through a 40µm nylon mesh. Purify protoplasts by centrifugation in a W5 solution sucrose cushion.
- Co-transformation: For each sample, mix 10 µg pUbi-Cas9D10A-Nickase plasmid, 10 µg of each pU6-gRNA plasmid, and 100 µL of protoplasts (2x10⁵ cells). Add 110 µL PEG-Calcium solution, mix gently, and incubate for 15 min at room temperature.
- Washing and Incubation: Slowly add 1 mL W5 solution, dilute, and pellet protoplasts. Resuspend in 1 mL culture medium. Incubate in the dark at 25°C for 48-72 hours.
- DNA Extraction and Analysis: Harvest protoplasts, extract genomic DNA. Perform targeted deep sequencing on the on-target and predicted off-target sites. Specificity is achieved via the generation of paired nicks; off-target sites lacking both paired gRNA matches will largely remain unmodified.
Diagrams
Title: Workflow for Designing & Testing CRISPR Specificity
Title: Strategic Categories to Minimize Off-Target Effects
1. Introduction & Thesis Context Within the broader thesis on Agrobacterium-mediated CRISPR-Cas9 delivery for functional genomics in recalcitrant plant species, a critical bottleneck is the regeneration of transformed, gene-edited plants. This protocol details a systematic approach to optimize regeneration media components and selection marker regimes for "difficult-to-transform" species (e.g., many monocots, legumes, and woody perennials). Success in this stage is paramount for obtaining non-chimeric, edited plants for subsequent phenotypic and molecular analysis in drug discovery pathways.
2. Quantitative Data Summary
Table 1: Comparative Efficacy of Cytokinin Types on Shoot Induction in Recalcitrant Species
| Cytokinin Type | Example Species | Concentration Range (µM) | Average Shoot Number (per explant) | Callus Formation Index (0-5) | Best Used For |
|---|---|---|---|---|---|
| 6-Benzylaminopurine (BAP) | Oryza sativa, Populus spp. | 0.5 - 5.0 | 2.3 ± 0.8 | 4 (High) | Species with high cytokinin tolerance |
| Thidiazuron (TDZ) | Glycine max, Malus domestica | 0.1 - 2.0 | 3.5 ± 1.2 | 5 (Very High) | Stimulating shoot organogenesis from callus |
| Zeatin | Solanum tuberosum, Pinus radiata | 0.5 - 10.0 | 1.8 ± 0.6 | 2 (Low) | Direct shoot organogenesis, minimizing callus |
| Kinetin | Cicer arietinum | 1.0 - 10.0 | 1.2 ± 0.5 | 3 (Medium) | Auxin-synergistic responses |
Table 2: Selection Marker Systems for CRISPR-Transformed Tissues
| Selection Marker | Mode of Action | Working Concentration | Average Escape Rate (%) | Advantage for Difficult Species |
|---|---|---|---|---|
| Hygromycin B phosphotransferase (hptII) | Inhibits protein synthesis | 10-50 mg/L (plant) | 15-25 | Broad efficacy, well-characterized |
| Neomycin phosphotransferase II (nptII) | Inhibits protein synthesis | 50-100 mg/L (kanamycin) | 20-30 | Low cost, common in dicots |
| Phosphinothricin acetyltransferase (pat/bar) | Inhibits glutamine synthetase | 2-10 mg/L (glufosinate) | 5-15 | Effective for monocots; chemical can also be sprayed |
| 2-Deoxyglucose-6-phosphate phosphatase (DOGR1) | Toxic sugar analog resistance | 10-20 mM (2-deoxyglucose) | <10 | Positive selection; no hormone interference |
3. Detailed Experimental Protocols
Protocol 3.1: Systematic Optimization of Plant Growth Regulators (PGRs) Objective: Identify the optimal type and concentration of auxins and cytokinins for callus induction and shoot regeneration from CRISPR-transformed explants. Materials: Sterile explants (e.g., immature embryos, leaf disks), basal medium (MS, B5), stock solutions of auxins (2,4-D, NAA, IAA) and cytokinins (BAP, TDZ, Zeatin), phytagel, sterile culture vessels. Procedure:
- Prepare a matrix of media with varying auxin (0.1-5.0 mg/L) and cytokinin (0-5.0 mg/L) concentrations.
- Inoculate 20 explants per treatment. Maintain at 25°C under a 16/8h photoperiod.
- At 14-day intervals for 8 weeks, score for:
- Callus Induction Frequency (%): (No. of explants with callus / Total) * 100.
- Callus Morphology: Record as friable, compact, or embryogenic.
- Shoot Initiation Time (days) and Number of Shoots per Responding Explant.
- Transfer developed shoots (>2 cm) to a PGR-free rooting medium. Statistical analysis (ANOVA) identifies significant optimal combinations.
Protocol 3.2: A Dual-Selection Strategy for CRISPR-Edited Events Objective: Efficiently select stably transformed, edited cells while suppressing Agrobacterium overgrowth. Materials: Agrobacterium strain harboring CRISPR-T-DNA with hptII and nptII markers, explants, co-cultivation media, selection media containing hygromycin and cefotaxime/timentin. Procedure:
- Co-cultivation & Initial Recovery: Co-cultivate explants with Agrobacterium for 3 days. Transfer to recovery media with a bactericidal antibiotic (e.g., 500 mg/L cefotaxime) but no selection agent for 5 days.
- Dual Selection Phase: Transfer explants to selection media containing both:
- Hygromycin (25 mg/L): For primary selection of transformed plant cells.
- Carbenicillin (500 mg/L) or Timentin (300 mg/L): For continued Agrobacterium suppression, which can metabolize selection agents.
- Sub-culturing: Sub-culture surviving calli or tissues onto fresh dual-selection media every 14 days for 8 weeks.
- PCR Validation: Isolate genomic DNA from putative transgenic tissues. Perform PCR with primers for the hptII gene and the CRISPR-Cas9 target site to confirm integration and editing.
4. Visualizations
Diagram 1: Workflow for Optimizing Regeneration Post-CRISPR Delivery
Diagram 2: Key Nodes in Plant Regeneration Signaling Pathways
5. The Scientist's Toolkit: Key Research Reagent Solutions
| Item | Function & Application in Protocol |
|---|---|
| Thidiazuron (TDZ) | Potent cytokinin-like regulator; crucial for inducing shoot organogenesis in recalcitrant species like soybean and apple. |
| Zeatin (Trans-isomer) | Natural cytokinin; promotes direct shoot development with minimal callus, ideal for conifers and some monocots. |
| Hygromycin B | Aminoglycoside antibiotic; selective agent for plant transformation (via hptII gene); effective in both monocots and dicots. |
| Glufosinate (Basta) | Herbicide; used for selection with pat/bar genes; allows for spray assay on putative transgenic plants. |
| Timentin (Ticarcillin/Clavulanate) | β-lactam antibiotic combination; superior to carbenicillin/cefotaxime for suppressing persistent Agrobacterium. |
| Phytagel | Gelling agent; provides clear, firm medium superior to agar for visualizing and isolating transformed callus events. |
| Silver Nitrate (AgNO₃) | Ethylene inhibitor; added to regeneration media (1-10 mg/L) to improve shoot elongation in many species. |
| L-Proline | Osmoprotectant/amino acid; added at 50-200 mg/L to callus induction media to enhance somatic embryogenesis. |
Benchmarking Success: Validating Edits and Comparing Delivery Methods
Within the context of a thesis on Agrobacterium-mediated CRISPR-Cas9 delivery for plant genome editing, confirming the presence and sequence of intended edits is paramount. Post-transformation, plant tissues are screened, and potential transgenic events must be rigorously validated at the molecular level. This application note details protocols for the initial PCR-based detection of transgene integration, followed by comprehensive sequencing strategies to verify on-target edits and assess editing efficiency.
PCR-Based Screening for Transgene Presence
Protocol: Genomic DNA Extraction from Plant Leaf Tissue (CTAB Method)
Materials:
- 100 mg fresh leaf tissue.
- 1.5 mL microcentrifuge tubes.
- Liquid Nitrogen.
- CTAB Extraction Buffer (pre-warmed to 65°C): 2% CTAB, 100 mM Tris-HCl (pH 8.0), 20 mM EDTA, 1.4 M NaCl, 1% PVP-40.
- Chloroform:Isoamyl Alcohol (24:1).
- Isopropanol.
- 70% Ethanol.
- TE Buffer or Nuclease-free Water.
- Mortar and pestle or tissue lyser.
Procedure:
- Grind ~100 mg leaf tissue to a fine powder in liquid nitrogen.
- Transfer powder to a tube with 700 µL pre-warmed CTAB buffer. Mix thoroughly.
- Incubate at 65°C for 30-60 minutes, inverting gently every 10 minutes.
- Cool to room temperature. Add 700 µL chloroform:isoamyl alcohol. Mix by inversion for 10 minutes.
- Centrifuge at >12,000 g for 15 minutes at room temperature.
- Transfer the upper aqueous phase to a new tube. Add 0.7 volumes of isopropanol. Mix by inversion.
- Centrifuge at >12,000 g for 10 minutes to pellet DNA.
- Discard supernatant. Wash pellet with 500 µL 70% ethanol. Centrifuge for 5 minutes.
- Air-dry pellet for 10-15 minutes. Resuspend in 50-100 µL TE buffer or water.
- Quantify DNA using a spectrophotometer.
Protocol: End-Point PCR for Cas9/Guide RNA Cassette Detection
Reagents:
- Extracted genomic DNA (template, 50-100 ng).
- PCR Master Mix (includes Taq polymerase, dNTPs, MgCl₂, buffer).
- Forward and Reverse primers specific to the Cas9 gene or plant selection marker.
- Nuclease-free water.
Primer Design: Primers should amplify a 150-800 bp fragment internal to the integrated T-DNA.
Thermocycler Program:
| Step | Temperature | Time | Cycles |
|---|---|---|---|
| Initial Denaturation | 94-95°C | 3-5 min | 1 |
| Denaturation | 94-95°C | 30 sec | |
| Annealing | 55-65°C* | 30 sec | 30-35 |
| Extension | 72°C | 1 min/kb | |
| Final Extension | 72°C | 5-10 min | 1 |
| Hold | 4-10°C | ∞ | 1 |
Note: Annealing temperature is primer-specific.
Analysis: Run PCR products on a 1-2% agarose gel. A band of expected size indicates potential transgenic events for further analysis.
Research Reagent Solutions for PCR Screening
| Item | Function | Example/Note |
|---|---|---|
| CTAB Buffer | Lyses plant cells, denatures proteins, and complexes with polysaccharides to purify DNA. | Critical for polysaccharide-rich plant tissue. |
| Taq DNA Polymerase | Thermostable enzyme for amplifying target DNA sequences via PCR. | Standard for endpoint detection. |
| dNTP Mix | Provides nucleotides (A, T, C, G) for DNA strand synthesis during PCR. | |
| Sequence-Specific Primers | Short oligonucleotides that define the start and end points of amplification. | Must be designed for the specific transgene. |
| DNA Ladder | A molecular weight marker to estimate the size of PCR amplicons on a gel. | Essential for gel analysis. |
| Gel Stain (e.g., Ethidium Bromide, SYBR Safe) | Intercalates with DNA to allow visualization under UV light. | SYBR Safe is a safer alternative. |
Sequencing-Based Validation of On-Target Edits
PCR-positive events require sequencing to confirm the precise nature of the edit at the target genomic locus.
Protocol: Sanger Sequencing of Cloned PCR Amplicons
This gold-standard method distinguishes between mono- and bi-allelic edits in primary transformants.
Procedure:
- Amplify Target Locus: Perform PCR (as in 1.2) using primers flanking the CRISPR target site, using a high-fidelity polymerase.
- Gel Purify Amplicon: Isolate the specific band from an agarose gel and purify using a commercial kit.
- Clone Amplicon: Ligate the purified PCR product into a T/A or blunt-end cloning vector. Transform into competent E. coli.
- Colony PCR: Screen bacterial colonies by PCR to identify those with an insert.
- Plasmid Preparation: Culture positive colonies, isolate plasmid DNA.
- Sanger Sequencing: Sequence 8-16 individual clones per plant event using vector-specific or insert-specific primers.
- Analysis: Align sequences to the wild-type reference to identify insertions, deletions (indels), or substitutions.
Quantitative Data - Editing Efficiency Calculation: Editing efficiency can be quantified from clonal sequencing data.
Table 1: Example Calculation of Editing Efficiency from Clonal Sequencing
| Plant Event # | Total Clones Sequenced | Clones with Indel | Clones Wild-Type | Editing Efficiency |
|---|---|---|---|---|
| Plant 1 | 16 | 12 | 4 | 75% |
| Plant 2 | 12 | 3 | 9 | 25% |
| Plant 3 | 10 | 0 | 10 | 0% |
Editing Efficiency = (Number of clones with indels / Total clones sequenced) x 100
Protocol: Next-Generation Sequencing (NGS) for Population Analysis
NGS provides deep, quantitative analysis of editing outcomes in a pooled sample, ideal for assessing mutagenesis frequency and spectrum.
Procedure:
- Amplify Target Loci: For multiple targets, perform a multiplex PCR to amplify all genomic regions of interest from pooled plant DNA. Add sample-specific barcodes in a second PCR round.
- Library Purification & Quantification: Purify amplicons and quantify precisely (e.g., via fluorometry).
- Pool Libraries: Combine equimolar amounts of each barcoded library.
- Sequencing: Run on an NGS platform (e.g., Illumina MiSeq) to obtain >10,000 reads per target per sample.
- Bioinformatic Analysis: Use specialized tools (e.g., CRISPResso2, Cas-Analyzer) to align reads to a reference and quantify indels and homology-directed repair (HDR) events.
Research Reagent Solutions for Sequencing Validation
| Item | Function | Example/Note |
|---|---|---|
| High-Fidelity Polymerase | PCR enzyme with proofreading for accurate amplification prior to sequencing. | e.g., Pfu, Q5. Reduces PCR errors. |
| PCR Cloning Kit | Vector and competent cells for ligation and propagation of PCR amplicons. | e.g., pGEM-T Easy, TOPO TA kits. |
| Sanger Sequencing Service | Provides capillary electrophoresis-based DNA sequencing. | Outsourced to core facilities. |
| NGS Amplicon Library Prep Kit | Reagents for attaching sequencing adapters and barcodes to target amplicons. | Essential for Illumina workflows. |
| CRISPR NGS Analysis Software | Bioinformatics tool to process sequencing reads and quantify editing outcomes. | e.g., CRISPResso2, TIDE, ICE. |
Workflow & Analysis Diagrams
Title: Molecular Validation Workflow for CRISPR Edits
Title: Sanger Sequencing Analysis of Cloned Alleles
Phenotypic Screening and Confirmation of Biallelic/Monoallelic Mutations
1. Introduction and Context within Agrobacterium-mediated CRISPR Research
The deployment of Agrobacterium tumefaciens for CRISPR-Cas9 delivery into plant cells is a cornerstone of modern plant functional genomics and trait development. Unlike biolistic methods, T-DNA integration often results in complex, multicopy insertion events. This necessitates robust downstream frameworks for phenotypic screening and genotypic confirmation of mutation zygosity (biallelic vs. monoallelic). This protocol details an integrated workflow, from initial screening of transgenic events to precise molecular characterization, critical for validating gene function and advancing edited lines toward commercial development.
2. Phenotypic Screening Workflow for Putative Mutants
Initial screening leverages selectable markers and visible phenotypes to enrich for potential knockout events before resource-intensive genotyping.
Protocol 2.1: Primary Selection and Visual Phenotype Assessment
- Materials: T0 seeds harvested from Agrobacterium-infiltrated plants, appropriate selective antibiotic or herbicide media, growth chambers.
- Method:
- Surface-sterilize T0 seeds and plate onto solid MS medium containing the selection agent (e.g., hygromycin, kanamycin, or glufosinate) corresponding to the T-DNA vector.
- Incubate under standard growth conditions for 2-3 weeks.
- Identify and separate resistant (putative transgenic) seedlings from sensitive (non-transgenic) seedlings.
- For edits targeting a known visual trait (e.g., albinism, dwarfism, altered leaf morphology), transfer resistant seedlings to soil and monitor for the expected phenotype over subsequent growth stages.
- Documentation: Record survival rates on selection and the incidence of any visible phenotypic deviations.
Table 1: Example Phenotypic Screening Data from a Chlorophyll Deficiency Edit
| T0 Line | Seeds Screened | Resistant Seedlings | Selection Rate (%) | Visible Mutant Phenotype (e.g., Albino) | Phenotypic Penetrance (%) |
|---|---|---|---|---|---|
| Line_01 | 120 | 34 | 28.3 | 8 | 23.5 |
| Line_02 | 115 | 41 | 35.7 | 0 | 0.0 |
| Line_03 | 122 | 29 | 23.8 | 11 | 37.9 |
3. Genotypic Confirmation of Mutations
Phenotypic prescreening is followed by DNA-level analysis to confirm CRISPR-induced edits and determine their zygosity.
Protocol 3.1: DNA Extraction and Target Region Amplification
- Materials: Leaf tissue, genomic DNA extraction kit (e.g., CTAB method), high-fidelity PCR polymerase, target-specific primers flanking the CRISPR-Cas9 cut site.
- Method:
- Isolate genomic DNA from ~100 mg of leaf tissue from selected T0 plants.
- Design primers ~300-500bp upstream and downstream of the intended cut site. Verify specificity in silico.
- Perform PCR amplification under optimized, high-fidelity conditions.
- Analyze PCR products on a 1% agarose gel for a single amplicon of expected size.
- Purify PCR products for downstream sequencing.
Protocol 3.2: Sanger Sequencing and Trace Deconvolution
- Materials: Purified PCR product, Sanger sequencing service, sequence analysis software (e.g., SnapGene, ICE Analysis [Synthego], TIDE).
- Method:
- Submit purified PCR amplicons for Sanger sequencing from both forward and reverse directions.
- Import sequencing trace (.ab1) files into analysis software.
- For Wild-type/Monoallelic Mutations: Align sequence to the reference. A clean, single-sequence trace after the cut site indicates a wild-type or homozygous edit. Overlapping peaks starting at the cut site indicate a monoallelic mutation (one allele edited, one wild-type).
- For Biallelic/Complex Mutations: Use deconvolution tools (ICE or TIDE). Input the edited sample trace and a control wild-type trace. The software quantifies the mixture of indel sequences.
- Interpretation: The tool outputs a readout of inferred indel sequences and their percentages.
Table 2: Genotype Analysis Results from Sanger Trace Deconvolution
| Plant ID | Phenotype | Inferred Genotypes (from ICE Analysis) | Allele 1 Frequency | Allele 2 Frequency | Zygosity Call | |
|---|---|---|---|---|---|---|
| Line_03-08 | Strong Mutant | 1-bp deletion | 7-bp deletion | 48% | 46% | Biallelic Mutant |
| Line_01-12 | Weak Mutant | Wild-type sequence | 4-bp insertion | 58% | 42% | Monoallelic Mutant |
| Line_02-05 | Wild-type | Wild-type sequence | Wild-type sequence | ~100% | ~0% | Wild-type |
| Line_03-11 | Strong Mutant | Complex rearrangement (15bp replacement) | 2-bp deletion | 52% | 48% | Biallelic Mutant |
4. The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Materials for Phenotypic and Genotypic Screening
| Item Name | Function & Application |
|---|---|
| Agrobacterium Strain (e.g., EHA105, GV3101) | Delivery vector for CRISPR-T-DNA into plant cells. Strain choice impacts transformation efficiency. |
| Selective Agent (e.g., Hygromycin B, Glufosinate) | Kills non-transformed tissue in planta or in vitro, allowing growth only of T-DNA-containing events. |
| High-Fidelity PCR Polymerase (e.g., Q5, Phusion) | Accurately amplifies the target genomic locus from plant DNA without introducing errors. |
| Sanger Sequencing Service | Provides capillary electrophoresis traces for the PCR amplicon, the raw data for mutation detection. |
| Trace Deconvolution Software (ICE or TIDE) | Algorithmically decomposes complex Sanger traces into constituent indel sequences and their relative abundances. |
| Next-Generation Sequencing (NGS) Kit (for amplicon-seq) | Provides ultra-deep sequencing of target amplicons for quantifying editing efficiency and detecting rare edits in pooled samples. |
Within the context of developing robust Agrobacterium-mediated CRISPR delivery systems for plant genome editing, a critical initial choice is the transformation methodology. Two principal techniques dominate: Agrobacterium tumefaciens-mediated transformation (ATMT) and direct DNA delivery via biolistics (gene gun). This analysis compares their pros, cons, and protocols, providing a framework for selecting the optimal approach for CRISPR construct delivery in plant cells.
Comparative Analysis: Core Features
Table 1: Qualitative Comparison of Key Characteristics
| Characteristic | Agrobacterium-Mediated Transformation | Biolistics (Gene Gun) |
|---|---|---|
| Principle | Biological; uses bacterial vector to transfer T-DNA | Physical; uses kinetic energy to deliver coated particles |
| Typical Insert Size | Large (>50 kb possible) | Moderate (up to ~40 kb) |
| Copy Number | Usually low (1-3 copies) | Often high, complex integration patterns |
| Integration Fidelity | Precise, with defined right-border end | Less precise, frequent truncation/rearrangement |
| Host Range | Limited primarily to dicots and some monocots | Extremely broad; any cell type |
| Tissue Culture Requirement | Generally required for regeneration | Can be used on non-dividing cells & intact tissues |
| Cost & Equipment | Low; requires standard microbiology lab | High; requires particle gun device & consumables |
| Regulatory/Ethical Perception | Often viewed as a "natural" process | Considered a more direct, transgene-specific method |
Table 2: Quantitative Performance Metrics
| Metric | Agrobacterium | Biolistics | Notes |
|---|---|---|---|
| Transformation Efficiency (Model Plant) | 50-90% (stable, in amenable species) | 5-30% (transient); stable typically lower | Efficiency is highly species- and tissue-dependent. |
| Time to Regenerated Plant | 3-6 months | 6-9 months | Biolistics often requires longer selection due to complex integration. |
| Frequency of Single-Copy Events | High (70-80%) | Low (10-30%) | Crucial for simplifying molecular characterization. |
| Labor Intensity (Post-Delivery) | Moderate | High | Due to high copy number and complex patterns in biolistics. |
Detailed Protocols
Protocol 1:Agrobacterium-Mediated CRISPR Delivery to Plant Leaf Disks
This protocol is optimized for delivering a CRISPR/Cas9 construct to dicot species (e.g., tobacco, tomato, Arabidopsis).
I. Materials Preparation
- Plant Material: Sterile leaf disks from in vitro grown plants.
- Agrobacterium Strain: A. tumefaciens GV3101 pSoup, carrying binary vector with CRISPR/Cas9 and plant selection marker.
- Media:
- YEP Liquid Media (for Agrobacterium growth): 10 g/L peptone, 10 g/L yeast extract, 5 g/L NaCl, pH 7.0.
- Co-cultivation Media (CCM): MS basal salts, vitamins, 3% sucrose, 1 mg/L 6-BA, 0.1 mg/L NAA, 200 µM acetosyringone, 0.8% agar.
- Selection Media: CCM + appropriate antibiotic (e.g., Kanamycin) + Timentin (500 mg/L) to kill Agrobacterium.
- Solutions: 10% Sodium Hypochlorite (bleach), 0.1% HgCl₂ (or commercial sterilant), sterile distilled water, 200 mM acetosyringone stock (in DMSO).
II. Step-by-Step Procedure
- Vector Transformation: Introduce the CRISPR binary vector into Agrobacterium via electroporation or freeze-thaw. Select on YEP plates with appropriate antibiotics.
- Bacterial Culture: Inoculate a single colony into 5 mL YEP with antibiotics. Grow overnight (28°C, 200 rpm). Dilute 1:50 in fresh media and grow to OD₆₀₀ ≈ 0.6-0.8.
- Induction: Pellet cells. Resuspend in liquid CCM (without agar) to OD₆₀₀ ≈ 0.5. Add acetosyringone to 200 µM. Incubate at 28°C, 100 rpm for 2-4 hours.
- Plant Explant Preparation: Surface sterilize leaves (10% bleach, 10 min), rinse 3x with sterile water. Punch 5-10 mm disks.
- Co-cultivation: Immerse leaf disks in the induced Agrobacterium suspension for 10-20 minutes. Blot dry on sterile paper. Place disks abaxial side down on solid CCM. Co-cultivate in the dark at 22-25°C for 48-72 hours.
- Selection & Regeneration: Transfer disks to Selection Media. Subculture to fresh media every 2 weeks. Shoot primordia appear in 3-6 weeks.
- Rooting & Molecular Analysis: Excise shoots and transfer to rooting medium. Confirm CRISPR editing via PCR/RE assay and sequencing of target loci in regenerated plants.
Protocol 2: Biolistic Delivery of CRISPR Ribonucleoproteins (RNPs) to Embryogenic Callus
This protocol emphasizes the delivery of pre-assembled Cas9-gRNA ribonucleoprotein complexes to minimize DNA integration, suitable for recalcitrant monocots.
I. Materials Preparation
- Plant Material: Embryogenic callus (e.g., from rice or maize immature embryos).
- Biolistics Device: PDS-1000/He or equivalent.
- Microcarriers: 0.6 µm gold or tungsten particles.
- Coating DNA/RNPs:
- For DNA: 5-10 µg purified CRISPR plasmid DNA.
- For RNP: 10 µg purified Cas9 protein pre-complexed with 5 µg in vitro transcribed sgRNA (30 min, 25°C).
- Solutions: 2.5 M CaCl₂, 0.1 M spermidine (free base, sterile), absolute ethanol, 70% ethanol, sterile TE buffer (pH 8.0).
- Media: Osmoticum treatment media (culture media + 0.2-0.4 M sorbitol/mannitol), standard recovery & selection media.
II. Step-by-Step Procedure
- Microcarrier Preparation: Weigh 60 mg of gold particles into a 1.5 mL tube. Add 1 mL 70% ethanol, vortex, incubate 15 min. Pellet, wash 3x with sterile water, then 1x with 50% glycerol. Resuspend in 1 mL 50% glycerol. Store at -20°C.
- Coating Microcarriers (for DNA):
- Aliquot 50 µL washed particles (in water or ethanol) to a new tube.
- Sequentially add (with continuous vortexing): 5 µL DNA (1 µg/µL), 50 µL 2.5 M CaCl₂, 20 µL 0.1 M spermidine.
- Vortex 2-3 min. Let settle 1 min. Pellet briefly. Remove supernatant.
- Wash with 140 µL 70% ethanol, then 140 µL 100% ethanol. Resuspend in 48 µL 100% ethanol.
- Coating Microcarriers (for RNP): Follow a similar precipitation protocol, but use NLS-tagged Cas9-sgRNA RNP complex instead of DNA, and use PEG instead of spermidine to aid precipitation.
- Target Preparation: Place embryogenic callus (fresh or osmoticum-treated for 4 hours) in the center of a petri dish containing osmotic media. Place dish in the bombardment chamber at the appropriate stopping distance (typically 6 or 9 cm).
- Bombardment: Follow device-specific instructions. For PDS-1000/He: Use 1100 psi rupture discs, vacuum of 28 in Hg, fire.
- Post-Bombardment Recovery: After bombardment, incubate tissues on osmotic media for 16-24 hours. Transfer to standard recovery media for 1 week, then to selection media if using DNA delivery.
- Analysis: For RNP delivery, analyze callus cells for editing within 48-72 hours via T7E1 or next-gen sequencing, as no stable integration is expected. For DNA, proceed with selection and regeneration as per standard protocols.
Visualizations
(Pathway: Agrobacterium T-DNA Transfer Mechanism)
(Workflow: Agrobacterium CRISPR Plant Transformation)
(Workflow: Biolistics CRISPR Delivery Pathways)
The Scientist's Toolkit
Table 3: Essential Research Reagent Solutions
| Item | Function in Protocol | Key Considerations |
|---|---|---|
| Binary Vector System (e.g., pCAMBIA, pGreen) | Carries CRISPR/Cas9 expression cassette within T-DNA borders for Agrobacterium delivery. | Choose based on plant selection marker, reporter genes, and replication origin compatibility with Agrobacterium helper strain. |
| Agrobacterium Helper Strain (e.g., GV3101, LBA4404, EHA105) | Provides Virulence (vir) genes in trans to mobilize T-DNA from the binary vector into the plant cell. | Strain choice affects host range and transformation efficiency. Disarmed (non-oncogenic) strains are standard. |
| Acetosyringone | A phenolic compound that activates the Agrobacterium vir gene expression system, mimicking plant wound signals. | Critical for transforming many plant species, especially monocots. Use fresh stock solution in DMSO. |
| Gold Microcarriers (0.6-1.0 µm) | Inert particles that serve as the physical carrier for DNA or RNPs during biolistic bombardment. | Size affects penetration and damage. Gold is chemically inert and non-toxic compared to tungsten. |
| Purified Cas9 Protein (for RNP Delivery) | The CRISPR nuclease enzyme, pre-complexed with sgRNA to form an active editing complex. | Enables transient, DNA-free editing. Reduces off-target integration. Requires optimization of NLS tags and purity. |
| Osmoticum (Sorbitol/Mannitol) | Added to plant tissue media pre-/post-bombardment to plasmolyze cells, reducing turgor pressure and cell damage from particle impact. | Increases transient expression and stable transformation efficiency in biolistics. Concentration is species-specific. |
| Timentin (or Carbenicillin) | Antibiotic used in plant culture media to eliminate residual Agrobacterium after co-cultivation. | Prevents overgrowth of Agrobacterium on plant tissues. Less phytotoxic than other beta-lactams like carbenicillin for some species. |
Within the framework of advancing Agrobacterium-mediated CRISPR delivery for plant genome engineering, novel delivery vectors have emerged. The following application notes compare the established Agrobacterium tumefaciens system with nanoparticle-based and viral vector delivery, summarizing key quantitative metrics.
Table 1: Comparative Analysis of Plant Cell Delivery Methods
| Parameter | Agrobacterium tumefaciens | Nanoparticles (e.g., Carbon Nanotubes, PEI) | Viral Vectors (e.g., Tobacco Rattle Virus, Bean Yellow Dwarf Virus) |
|---|---|---|---|
| Max Cargo Capacity | >50 kbp (T-DNA, whole genes) | ~2-20 kbp (plasmid DNA) | 1.5 - 10 kbp (viral genome dependent) |
| Delivery Efficiency (Transient) | Moderate to High (species-dependent) | Low to Moderate (highly variable) | Very High (systemic infection) |
| Stable Integration Rate | High (T-DNA integration) | Extremely Low (mostly transient) | Extremely Low (non-integrative) |
| Host Range | Primarily dicots; some monocots | Very Broad (physical delivery) | Narrow to Moderate (host-specific) |
| Throughput & Scalability | Moderate (bacterial culture required) | High (protocols facile) | High (inoculation simple) |
| Regulatory Considerations | GMO regulations apply | Emerging regulatory path | GMO regulations apply |
Detailed Application Notes & Protocols
Agrobacterium tumefaciens(GV3101) Mediated CRISPR Delivery
- Principle: Utilizes the natural Ti-plasmid machinery to transfer T-DNA containing CRISPR-Cas9 expression cassettes into the plant cell nucleus for stable integration or transient expression.
- Key Application: Generation of stable, heritable gene-edited lines.
Protocol A: Stable Transformation of Nicotiana benthamiana Leaf Discs
- Day 1: Culture Preparation. Inoculate 5 mL of YEP (with appropriate antibiotics for Agrobacterium strain and binary vector) with a single colony of A. tumefaciens GV3101 harboring the CRISPR plasmid. Grow overnight (28°C, 250 rpm).
- Day 2: Infection. Dilute the overnight culture to OD₆₀₀ = 0.5 in liquid co-cultivation medium (MS salts, 3% sucrose, 200 µM acetosyringone). Aseptically subculture leaf discs into the bacterial suspension for 20 minutes. Blot discs on sterile paper and place on solid co-cultivation medium. Incubate in the dark (25°C) for 48-72 hours.
- Day 4-5: Selection & Regeneration. Transfer discs to selection/regeneration medium (containing appropriate antibiotics for plant selection, e.g., kanamycin or hygromycin, and 500 mg/L cefotaxime to kill Agrobacterium). Subculture every 2 weeks until shoot formation.
- Day 28+: Rooting & Acclimatization. Excise shoots and transfer to rooting medium. Once rooted, transfer plantlets to soil and acclimatize.
The Scientist's Toolkit: Key Reagents for Agrobacterium Protocol
| Reagent/Material | Function |
|---|---|
| A. tumefaciens GV3101 | Disarmed, helper plasmid-containing strain for efficient T-DNA transfer. |
| Binary Vector pCambia | Carries T-DNA with CRISPR-Cas9 and plant selection marker. |
| Acetosyringone | Phenolic compound that induces vir gene expression in Agrobacterium. |
| Cefotaxime | Beta-lactam antibiotic used to eliminate Agrobacterium post-co-cultivation. |
| Selection Antibiotic (e.g., Kanamycin) | Selects for plant cells that have integrated the T-DNA. |
Diagram Title: Agrobacterium CRISPR Workflow
Nanoparticle-Mediated CRISPR-Cas9 RNP Delivery
- Principle: Chemically or physically assembled nanoparticles (e.g., cationic polymers, mesoporous silica) complex with, protect, and deliver pre-assembled CRISPR-Cas9 ribonucleoproteins (RNPs) into plant cells via passive or active (e.g., peptide-guided) uptake mechanisms.
Protocol B: Polyethyleneimine (PEI)-Coated Mesoporous Silica Nanoparticle (MSN) Delivery to Protoplasts
- Step 1: RNP Complexation. Pre-assemble purified Cas9 protein with sgRNA (molar ratio 1:2) in nuclease-free buffer (30 min, RT). Mix 10 µg of MSNs (amine-functionalized, 50 nm) with 5 µg of RNPs in 100 µL MES buffer (pH 5.5). Add 1% (w/v) PEI solution dropwise. Incubate 30 min (RT) to form RNP-PEI-MSN complexes.
- Step 2: Plant Protoplast Preparation. Isolate protoplasts from target tissue (e.g., leaf mesophyll) using an enzymatic digestion solution (1.5% cellulase, 0.4% macerozyme in 0.4 M mannitol). Purify via flotation/sedimentation.
- Step 3: Transfection. Resuspend 10⁵ protoplasts in 500 µL of MMg solution (0.4 M mannitol, 15 mM MgCl₂). Add the RNP-PEI-MSN complexes. Incubate for 30 minutes at RT. Add 2 volumes of W5 solution (154 mM NaCl, 125 mM CaCl₂, 5 mM KCl) to stop transfection.
- Step 4: Analysis. Incubate protoplasts in culture medium for 24-48 hours. Extract genomic DNA and assess editing efficiency via T7 Endonuclease I assay or next-generation sequencing.
Diagram Title: Nanoparticle RNP Delivery Pathway
Viral Vector-Mediated CRISPR Delivery (TRV-based VIGE)
- Principle: Engineered plant RNA viruses (e.g., Tobacco Rattle Virus - TRV) are used to deliver sgRNA sequences systemically throughout the plant. The Cas9 protein is expressed from a stable transgenic line or co-delivered via a separate viral vector.
Protocol C: Virus-Induced Genome Editing (VIGE) using TRV in N. benthamiana
- Step 1: Vector Construction. Clone the target-specific sgRNA expression cassette into the TRV RNA2 vector (pYL156 derivative) between appropriate restriction sites.
- Step 2: Agrobacterium Inoculum Prep. Transform constructs (TRV1, TRV2-sgRNA) into A. tumefaciens GV3101. Grow cultures, resuspend to OD₆₀₀ = 1.0 in infiltration buffer (10 mM MES, 10 mM MgCl₂, 200 µM acetosyringone). Mix TRV1 and TRV2-sgRNA cultures 1:1.
- Step 3: Plant Infiltration. Use a needleless syringe to pressure-infiltrate the mixed culture into the abaxial side of leaves of a transgenic Cas9-expressing N. benthamiana plant at the 4-6 leaf stage.
- Step 4: Systemic Infection & Analysis. Maintain plants for 2-3 weeks, allowing systemic viral movement. Harvest newly emerged, non-infiltrated systemic leaves. Extract DNA and analyze editing efficiency at the target locus.
Diagram Title: Viral Vector (VIGE) Systemic Delivery
Conclusion
Agrobacterium-mediated delivery remains a cornerstone technique for precise, stable CRISPR genome editing in plants, offering unique advantages in heritability and complex vector delivery. Mastering the foundational biology, robust methodologies, and optimization strategies outlined here is critical for successful application in crop engineering and basic research. Future directions will focus on expanding host range, developing transgene-free editing systems (e.g., using developmental regulators), and integrating Agrobacterium with next-generation CRISPR tools like prime editing. This synergy will accelerate the development of climate-resilient, nutritious crops and advance our fundamental understanding of plant biology, with profound implications for global food security and sustainable agriculture.